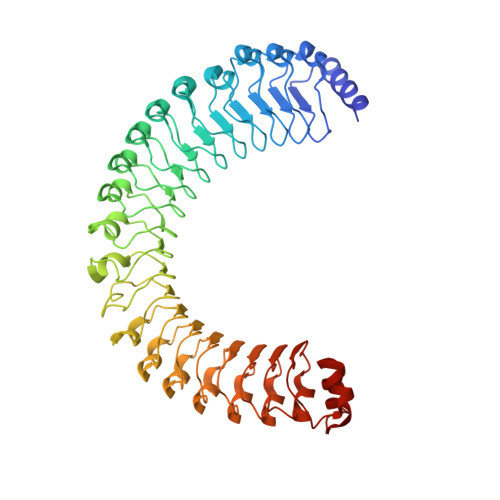
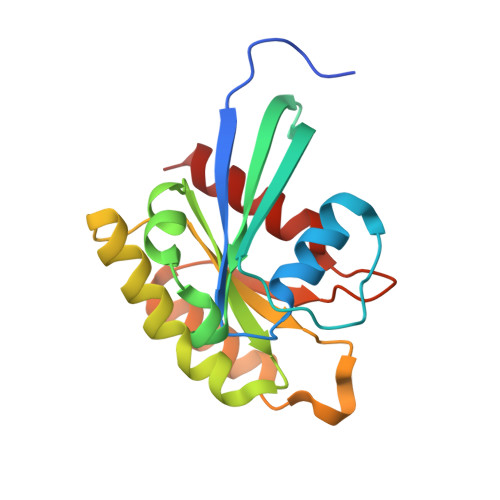
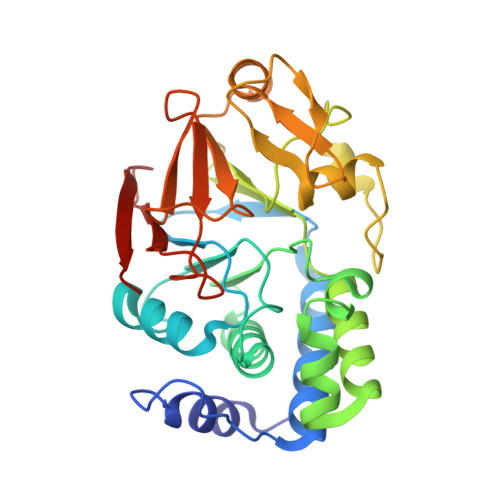
Structure of the MRAS-SHOC2-PP1C phosphatase complex.
Hauseman, Z.J., Fodor, M., Dhembi, A., Viscomi, J., Egli, D., Bleu, M., Katz, S., Park, E., Jang, D.M., Porter, K.A., Meili, F., Guo, H., Kerr, G., Molle, S., Velez-Vega, C., Beyer, K.S., Galli, G.G., Maira, S.M., Stams, T., Clark, K., Eck, M.J., Tordella, L., Thoma, C.R., King, D.A.(2022) Nature 609: 416-423
- PubMed: 35830882 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-022-05086-1
- Primary Citation Related Structures:
7TXH, 7TYG - PubMed Abstract:
RAS-MAPK signalling is fundamental for cell proliferation and is altered in most human cancers 1-3 . However, our mechanistic understanding of how RAS signals through RAF is still incomplete. Although studies revealed snapshots for autoinhibited and active RAF-MEK1-14-3-3 complexes 4 , the intermediate steps that lead to RAF activation remain unclear. The MRAS-SHOC2-PP1C holophosphatase dephosphorylates RAF at serine 259, resulting in the partial displacement of 14-3-3 and RAF-RAS association 3,5,6 . MRAS, SHOC2 and PP1C are mutated in rasopathies-developmental syndromes caused by aberrant MAPK pathway activation 6-14 -and SHOC2 itself has emerged as potential target in receptor tyrosine kinase (RTK)-RAS-driven tumours 15-18 . Despite its importance, structural understanding of the SHOC2 holophosphatase is lacking. Here we determine, using X-ray crystallography, the structure of the MRAS-SHOC2-PP1C complex. SHOC2 bridges PP1C and MRAS through its concave surface and enables reciprocal interactions between all three subunits. Biophysical characterization indicates a cooperative assembly driven by the MRAS GTP-bound active state, an observation that is extendible to other RAS isoforms. Our findings support the concept of a RAS-driven and multi-molecular model for RAF activation in which individual RAS-GTP molecules recruit RAF-14-3-3 and SHOC2-PP1C to produce downstream pathway activation. Importantly, we find that rasopathy and cancer mutations reside at protein-protein interfaces within the holophosphatase, resulting in enhanced affinities and function. Collectively, our findings shed light on a fundamental mechanism of RAS biology and on mechanisms of clinically observed enhanced RAS-MAPK signalling, therefore providing the structural basis for therapeutic interventions.
- Novartis Institutes for BioMedical Research, Cambridge, MA, USA. zachary.hauseman@novartis.com.
Organizational Affiliation: