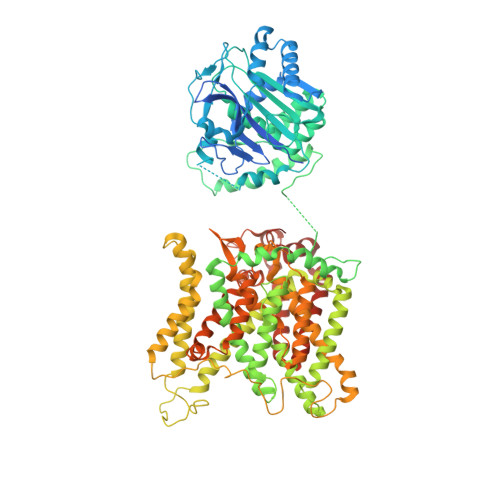
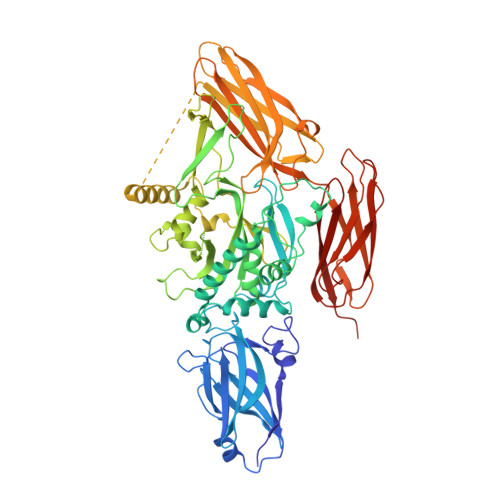
Structure, dynamics and assembly of the ankyrin complex on human red blood cell membrane.
Xia, X., Liu, S., Zhou, Z.H.(2022) Nat Struct Mol Biol 29: 698-705
- PubMed: 35655099 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-022-00779-7
- Primary Citation Related Structures:
7TVZ, 7TW0, 7TW1, 7TW2, 7TW3, 7TW5, 7TW6 - PubMed Abstract:
The cytoskeleton of a red blood cell (RBC) is anchored to the cell membrane by the ankyrin complex. This complex is assembled during RBC genesis and comprises primarily band 3, protein 4.2 and ankyrin, whose mutations contribute to numerous human inherited diseases. High-resolution structures of the ankyrin complex have been long sought-after to understand its assembly and disease-causing mutations. Here, we analyzed native complexes on the human RBC membrane by stepwise fractionation. Cryo-electron microscopy structures of nine band-3-associated complexes reveal that protein 4.2 stabilizes the cytoplasmic domain of band 3 dimer. In turn, the superhelix-shaped ankyrin binds to this protein 4.2 via ankyrin repeats (ARs) 6-13 and to another band 3 dimer via ARs 17-20, bridging two band 3 dimers in the ankyrin complex. Integration of these structures with both prior data and our biochemical data supports a model of ankyrin complex assembly during erythropoiesis and identifies interactions essential for the mechanical stability of RBC.
- Department of Microbiology, Immunology, and Molecular Genetics, University of California, Los Angeles, CA, USA.
Organizational Affiliation: