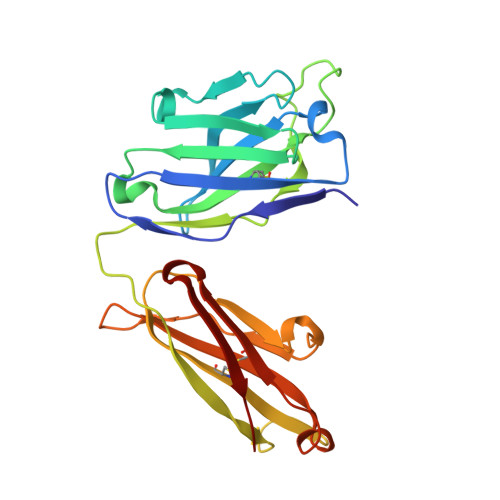
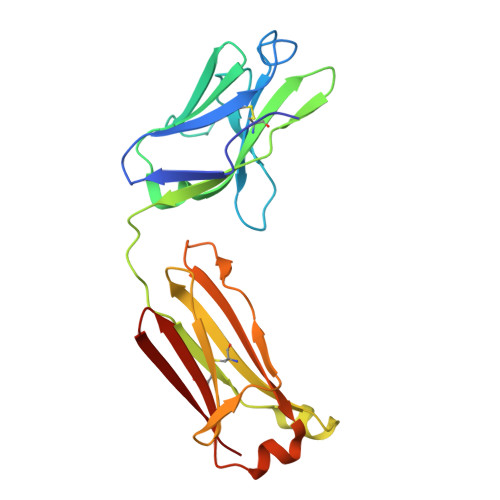
CD4 binding site immunogens elicit heterologous anti-HIV-1 neutralizing antibodies in transgenic and wild-type animals.
Gristick, H.B., Hartweger, H., Loewe, M., van Schooten, J., Ramos, V., Oliveira, T.Y., Nishimura, Y., Koranda, N.S., Wall, A., Yao, K.H., Poston, D., Gazumyan, A., Wiatr, M., Horning, M., Keeffe, J.R., Hoffmann, M.A.G., Yang, Z., Abernathy, M.E., Dam, K.A., Gao, H., Gnanapragasam, P.N.P., Kakutani, L.M., Pavlovitch-Bedzyk, A.J., Seaman, M.S., Howarth, M., McGuire, A.T., Stamatatos, L., Martin, M.A., West Jr., A.P., Nussenzweig, M.C., Bjorkman, P.J.(2023) Sci Immunol 8: eade6364-eade6364
- PubMed: 36763635
- DOI: https://doi.org/10.1126/sciimmunol.ade6364
- Primary Citation of Related Structures:
7TQG - PubMed Abstract:
Passive transfer of broadly neutralizing anti-HIV-1 antibodies (bNAbs) protects against infection, and therefore, eliciting bNAbs by vaccination is a major goal of HIV-1 vaccine efforts. bNAbs that target the CD4 binding site (CD4bs) on HIV-1 Env are among the most broadly active, but to date, responses elicited against this epitope in vaccinated animals have lacked potency and breadth. We hypothesized that CD4bs bNAbs resembling the antibody IOMA might be easier to elicit than other CD4bs antibodies that exhibit higher somatic mutation rates, a difficult-to-achieve mechanism to accommodate Env's N276 gp120 N-glycan, and rare five-residue light chain complementarity-determining region 3. As an initial test of this idea, we developed IOMA germline-targeting Env immunogens and evaluated a sequential immunization regimen in transgenic mice expressing germline-reverted IOMA. These mice developed CD4bs epitope-specific responses with heterologous neutralization, and cloned antibodies overcame neutralization roadblocks, including accommodating the N276 gp120 glycan, with some neutralizing selected HIV-1 strains more potently than IOMA. The immunization regimen also elicited CD4bs-specific responses in mice containing polyclonal antibody repertoires as well as rabbits and rhesus macaques. Thus, germline targeting of IOMA-class antibody precursors represents a potential vaccine strategy to induce CD4bs bNAbs.
- Division of Biology and Biological Engineering, California Institute of Technology, Pasadena, CA 91125, USA.
Organizational Affiliation: