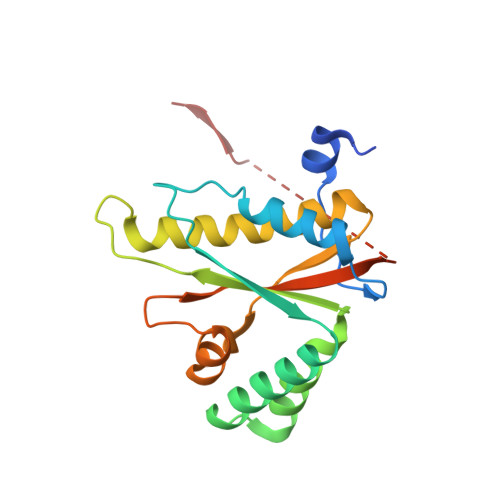
Crystal structures of NAD(P)H nitroreductases from Klebsiella pneumoniae
Kancherla, A.D., Liu, L., Tillery, L., Shek, R., Craig, J.K., Machen, A.J., Seibold, S., Battaile, K.P., Fradi, S., Barrett, L.K., Subramanian, S., Myler, P., Van Voorhis, W.C., Lovell, S.(2024) Acta Crystallogr F Struct Biol Commun F80: 173-182