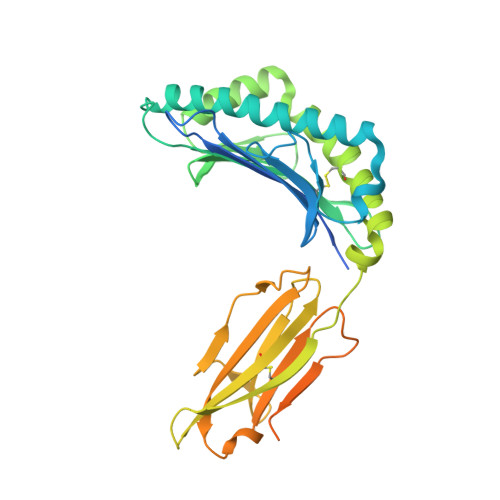
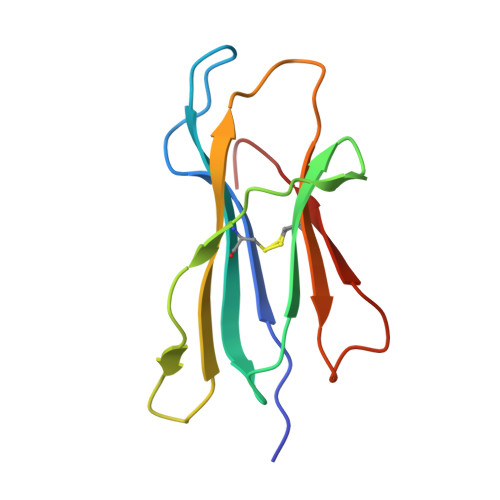
Ablation of CD8 + T cell recognition of an immunodominant epitope in SARS-CoV-2 Omicron variants BA.1, BA.2 and BA.3.
Swaminathan, S., Lineburg, K.E., Panikkar, A., Raju, J., Murdolo, L.D., Szeto, C., Crooks, P., Le Texier, L., Rehan, S., Dewar-Oldis, M.J., Barnard, P.J., Ambalathingal, G.R., Neller, M.A., Short, K.R., Gras, S., Khanna, R., Smith, C.(2022) Nat Commun 13: 6387-6387
- PubMed: 36302758 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-34180-1
- Primary Citation Related Structures:
7TLT - PubMed Abstract:
The emergence of the SARS-CoV-2 Omicron variant has raised concerns of escape from vaccine-induced immunity. A number of studies have demonstrated a reduction in antibody-mediated neutralization of the Omicron variant in vaccinated individuals. Preliminary observations have suggested that T cells are less likely to be affected by changes in Omicron. However, the complexity of human leukocyte antigen genetics and its impact upon immunodominant T cell epitope selection suggests that the maintenance of T cell immunity may not be universal. In this study, we describe the impact that changes in Omicron BA.1, BA.2 and BA.3 have on recognition by spike-specific T cells. These T cells constitute the immunodominant CD8 + T cell response in HLA-A*29:02 + COVID-19 convalescent and vaccinated individuals; however, they fail to recognize the Omicron-encoded sequence. These observations demonstrate that in addition to evasion of antibody-mediated immunity, changes in Omicron variants can also lead to evasion of recognition by immunodominant T cell responses.
- QIMR Berghofer Centre for Immunotherapy and Vaccine Development and Translational and Human Immunology Laboratory, Infection and Inflammation Program, QIMR Berghofer Medical Research Institute, Herston, QLD, 4006, Australia.
Organizational Affiliation: