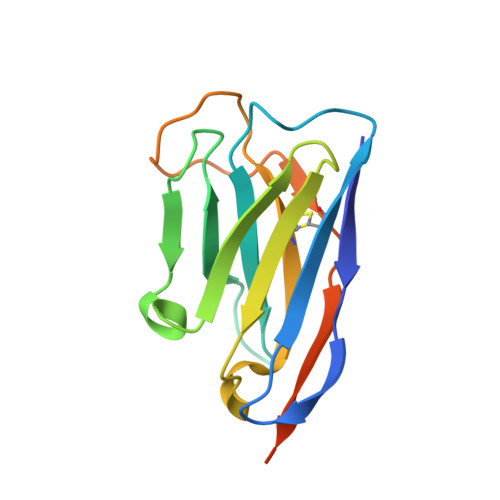
Structure and specificity of an anti-chloramphenicol single domain antibody for detection of amphenicol residues.
Swofford, C.A., Nordeen, S.A., Chen, L., Desai, M.M., Chen, J., Springs, S.L., Schwartz, T.U., Sinskey, A.J.(2022) Protein Sci 31: e4457-e4457
- PubMed: 36153664 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.4457
- Primary Citation Related Structures:
7TJC - PubMed Abstract:
Antibiotics in aquaculture prevent bacterial infection of fish, but their misuse is a public health risk and contributes to the unintentional creation of multiresistant pathogens. Regulatory agencies cannot do the rigorous, expensive testing required to keep up with the volume of seafood shipments. Current rapid test kits for these drugs enable the increase in testing needed for adequate monitoring of food supply chains, but they lack a high degree of accuracy. To combat this, we set out to discover and engineer single-domain antibodies (VHHs) that bind to small molecule antibiotics, and that can be used in rapid test kits. The small size, solubility, and stability of VHHs are useful properties that can improve the reliability and shelf-life of test kits for these adulterants. Here, we report a novel anti-chloramphenicol VHH (Chl-VHH) with a disassociation constant of 57 nM. This was achieved by immunizing a llama against a chloramphenicol-keyhole limpet hemocyanin (KLH) conjugate and screening for high affinity binders through phage display. The crystal structure of the bound-VHH to chloramphenicol was key to identifying a mutation in the binding pocket that resulted in a 16-fold improvement in binding affinity. In addition, the structure provides new insights into VHH-hapten interactions that can guide future engineering of VHHs against additional targets.
- Department of Biology, Massachusetts Institute of Technology, Cambridge, Massachusetts, USA.
Organizational Affiliation: