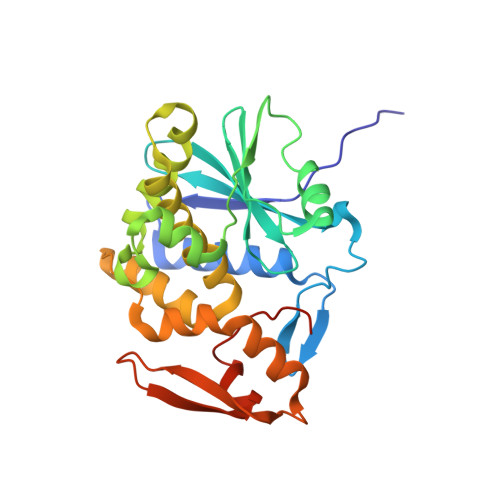
Single-domain antibodies neutralize ricin toxin intracellularly by blocking access to ribosomal P-stalk proteins.
Czajka, T.F., Vance, D.J., Davis, S., Rudolph, M.J., Mantis, N.J.(2022) J Biological Chem 298: 101742-101742
- PubMed: 35182523 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2022.101742
- Primary Citation Related Structures:
7TGF, 7TGI, 7TH2, 7TH3 - PubMed Abstract:
During ricin intoxication in mammalian cells, ricin's enzymatic (RTA) and binding (RTB) subunits disassociate in the endoplasmic reticulum. RTA is then translocated into the cytoplasm where, by virtue of its ability to depurinate a conserved residue within the sarcin-ricin loop (SRL) of 28S rRNA, it functions as a ribosome-inactivating protein. It has been proposed that recruitment of RTA to the SRL is facilitated by ribosomal P-stalk proteins, whose C-terminal domains interact with a cavity on RTA normally masked by RTB; however, evidence that this interaction is critical for RTA activity within cells is lacking. Here, we characterized a collection of single-domain antibodies (V H Hs) whose epitopes overlap with the P-stalk binding pocket on RTA. The crystal structures of three such V H Hs (V9E1, V9F9, and V9B2) in complex with RTA revealed not only occlusion of the ribosomal P-stalk binding pocket but also structural mimicry of C-terminal domain peptides by complementarity-determining region 3. In vitro assays confirmed that these V H Hs block RTA-P-stalk peptide interactions and protect ribosomes from depurination. Moreover, when expressed as "intrabodies," these V H Hs rendered cells resistant to ricin intoxication. One V H H (V9F6), whose epitope was structurally determined to be immediately adjacent to the P-stalk binding pocket, was unable to neutralize ricin within cells or protect ribosomes from RTA in vitro. These findings are consistent with the recruitment of RTA to the SRL by ribosomal P-stalk proteins as a requisite event in ricin-induced ribosome inactivation.
- Department of Biomedical Sciences, University at Albany, Albany, New York, USA.
Organizational Affiliation: