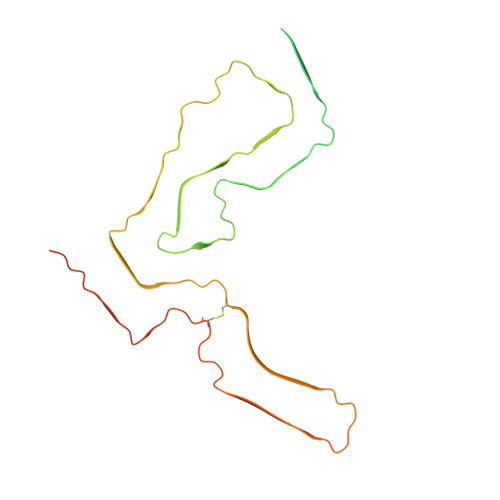
Cryo-EM structure of anchorless RML prion reveals variations in shared motifs between distinct strains.
Hoyt, F., Standke, H.G., Artikis, E., Schwartz, C.L., Hansen, B., Li, K., Hughson, A.G., Manca, M., Thomas, O.R., Raymond, G.J., Race, B., Baron, G.S., Caughey, B., Kraus, A.(2022) Nat Commun 13: 4005-4005
- PubMed: 35831291 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-30458-6
- Primary Citation Related Structures:
7TD6 - PubMed Abstract:
Little is known about the structural basis of prion strains. Here we provide a high (3.0 Å) resolution cryo-electron microscopy-based structure of infectious brain-derived fibrils of the mouse anchorless RML scrapie strain which, like the recently determined hamster 263K strain, has a parallel in-register β-sheet-based core. Several structural motifs are shared between these ex vivo prion strains, including an amino-proximal steric zipper and three β-arches. However, detailed comparisons reveal variations in these shared structural topologies and other features. Unlike 263K and wildtype RML prions, the anchorless RML prions lack glycophosphatidylinositol anchors and are severely deficient in N-linked glycans. Nonetheless, the similarity of our anchorless RML structure to one reported for wildtype RML prion fibrils in an accompanying paper indicates that these post-translational modifications do not substantially alter the amyloid core conformation. This work demonstrates both common and divergent structural features of prion strains at the near-atomic level.
- Research Technologies Branch, Rocky Mountain Laboratories, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Hamilton, MT, 59840, USA.
Organizational Affiliation: