Vaccine-elicited murine antibody WS6 neutralizes diverse beta-coronaviruses by recognizing a helical stem supersite of vulnerability.
Shi, W., Wang, L., Zhou, T., Sastry, M., Yang, E.S., Zhang, Y., Chen, M., Chen, X., Choe, M., Creanga, A., Leung, K., Olia, A.S., Pegu, A., Rawi, R., Schon, A., Shen, C.H., Stancofski, E.D., Talana, C.A., Teng, I.T., Wang, S., Corbett, K.S., Tsybovsky, Y., Mascola, J.R., Kwong, P.D.(2022) Structure 30: 1233
- PubMed: 35841885 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2022.06.004
- Primary Citation Related Structures:
7TCQ - PubMed Abstract:
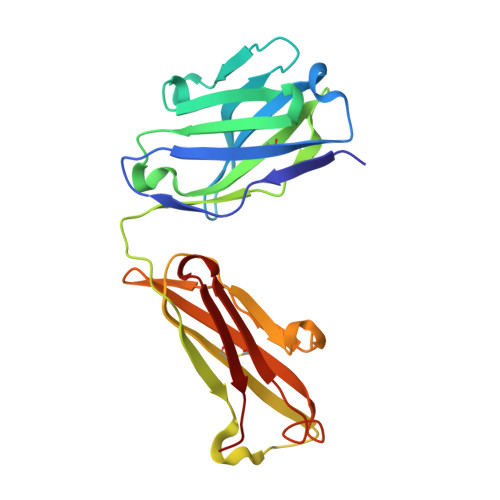
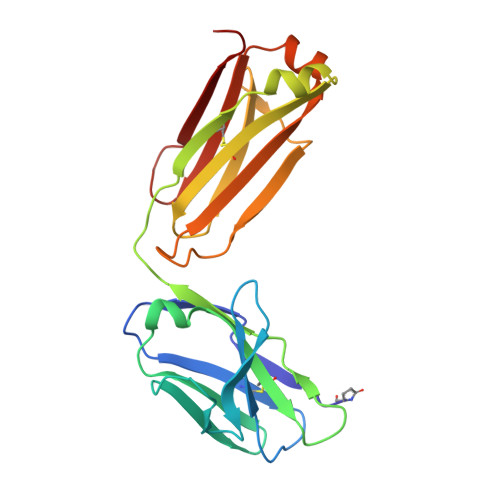
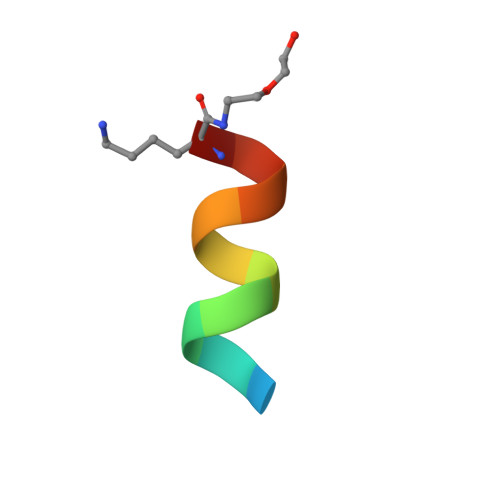
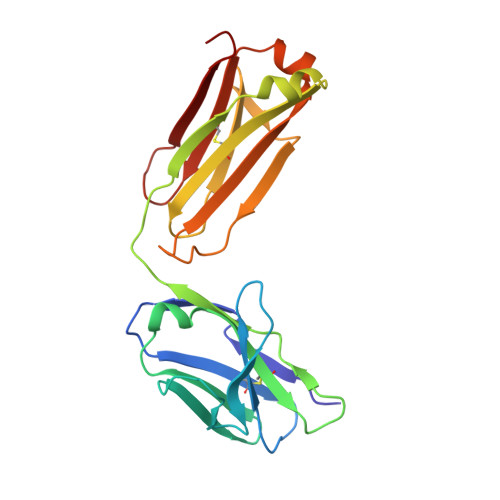
Immunization with severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) spike elicits diverse antibodies, but it is unclear if any of the antibodies can neutralize broadly against other beta-coronaviruses. Here, we report antibody WS6 from a mouse immunized with mRNA encoding the SARS-CoV-2 spike. WS6 bound diverse beta-coronavirus spikes and neutralized SARS-CoV-2 variants, SARS-CoV, and related sarbecoviruses. Epitope mapping revealed WS6 to target a region in the S2 subunit, which was conserved among SARS-CoV-2, Middle East respiratory syndrome (MERS)-CoV, and hCoV-OC43. The crystal structure at 2 Å resolution of WS6 revealed recognition to center on a conserved S2 helix, which was occluded in both pre- and post-fusion spike conformations. Structural and neutralization analyses indicated WS6 to neutralize by inhibiting fusion and post-viral attachment. Comparison of WS6 with other recently identified antibodies that broadly neutralize beta-coronaviruses indicated a stem-helical supersite-centered on hydrophobic residues Phe1148, Leu1152, Tyr1155, and Phe1156-to be a promising target for vaccine design.
- Vaccine Research Center, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD 20892, USA.
Organizational Affiliation: