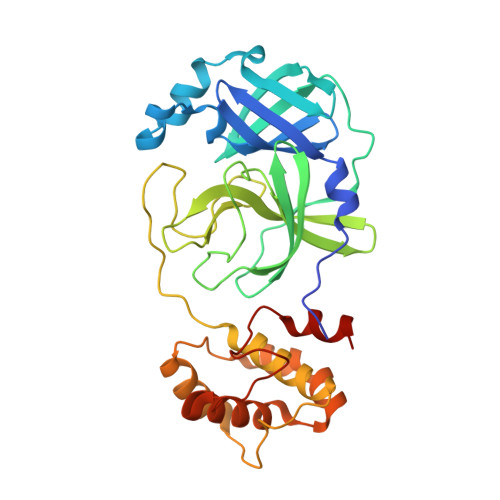
Structure-Guided Design of Potent Spirocyclic Inhibitors of Severe Acute Respiratory Syndrome Coronavirus-2 3C-like Protease.
Dampalla, C.S., Rathnayake, A.D., Galasiti Kankanamalage, A.C., Kim, Y., Perera, K.D., Nguyen, H.N., Miller, M.J., Madden, T.K., Picard, H.R., Thurman, H.A., Kashipathy, M.M., Liu, L., Battaile, K.P., Lovell, S., Chang, K.O., Groutas, W.C.(2022) J Med Chem 65: 7818-7832
- PubMed: 35638577 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.2c00224
- Primary Citation Related Structures:
7T3Y, 7T3Z, 7T40, 7T41, 7T42, 7T43, 7T44, 7T45, 7T46, 7T48, 7T49, 7T4A, 7T4B - PubMed Abstract:
The worldwide impact of the ongoing COVID-19 pandemic on public health has made imperative the discovery and development of direct-acting antivirals aimed at targeting viral and/or host targets. SARS-CoV-2 3C-like protease (3CL pro ) has emerged as a validated target for the discovery of SARS-CoV-2 therapeutics because of the pivotal role it plays in viral replication. We describe herein the structure-guided design of highly potent inhibitors of SARS-CoV-2 3CL pro that incorporate in their structure novel spirocyclic design elements aimed at optimizing potency by accessing new chemical space. Inhibitors of both SARS-CoV-2 3CL pro and MERS-CoV 3CL pro that exhibit nM potency and high safety indices have been identified. The mechanism of action of the inhibitors and the structural determinants associated with binding were established using high-resolution cocrystal structures.
- Department of Chemistry and Biochemistry, Wichita State University, Wichita, Kansas 67260, United States.
Organizational Affiliation: