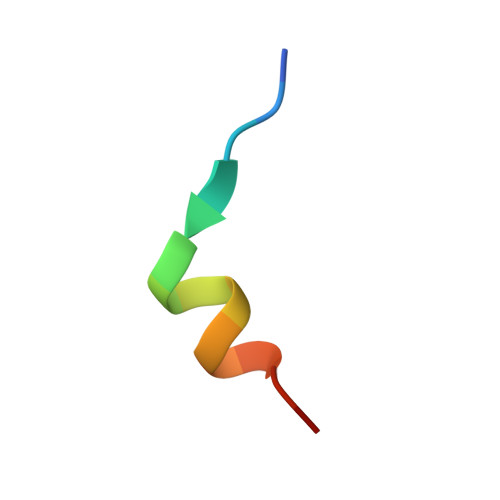
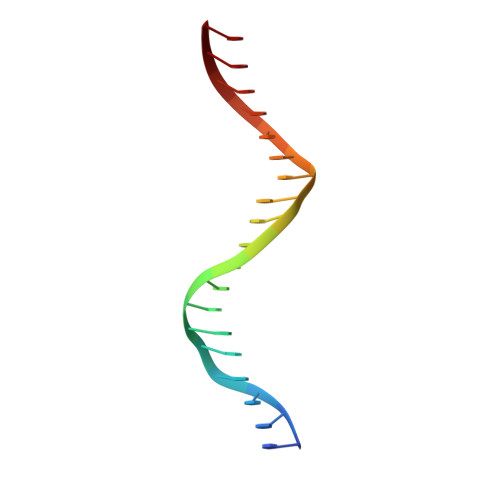
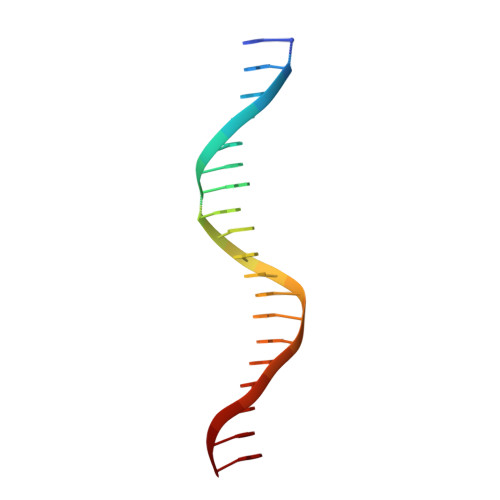
Mechanistic details of CRISPR-associated transposon recruitment and integration revealed by cryo-EM.
Park, J.U., Tsai, A.W., Chen, T.H., Peters, J.E., Kellogg, E.H.(2022) Proc Natl Acad Sci U S A 119: e2202590119-e2202590119
- PubMed: 35914146 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2202590119
- Primary Citation Related Structures:
7SVV, 7SVW - PubMed Abstract:
CRISPR-associated transposons (CASTs) are Tn7-like elements that are capable of RNA-guided DNA integration. Although structural data are known for nearly all core transposition components, the transposase component, TnsB, remains uncharacterized. Using cryo-electron microscopy (cryo-EM) structure determination, we reveal the conformation of TnsB during transposon integration for the type V-K CAST system from Scytonema hofmanni (ShCAST). Our structure of TnsB is a tetramer, revealing strong mechanistic relationships with the overall architecture of RNaseH transposases/integrases in general, and in particular the MuA transposase from bacteriophage Mu. However, key structural differences in the C-terminal domains indicate that TnsB's tetrameric architecture is stabilized by a different set of protein-protein interactions compared with MuA. We describe the base-specific interactions along the TnsB binding site, which explain how different CAST elements can function on cognate mobile elements independent of one another. We observe that melting of the 5' nontransferred strand of the transposon end is a structural feature stabilized by TnsB and furthermore is crucial for donor-DNA integration. Although not observed in the TnsB strand-transfer complex, the C-terminal end of TnsB serves a crucial role in transposase recruitment to the target site. The C-terminal end of TnsB adopts a short, structured 15-residue "hook" that decorates TnsC filaments. Unlike full-length TnsB, C-terminal fragments do not appear to stimulate filament disassembly using two different assays, suggesting that additional interactions between TnsB and TnsC are required for redistributing TnsC to appropriate targets. The structural information presented here will help guide future work in modifying these important systems as programmable gene integration tools.
- Department of Molecular Biology and Genetics, Cornell University, Ithaca, NY 14853.
Organizational Affiliation: