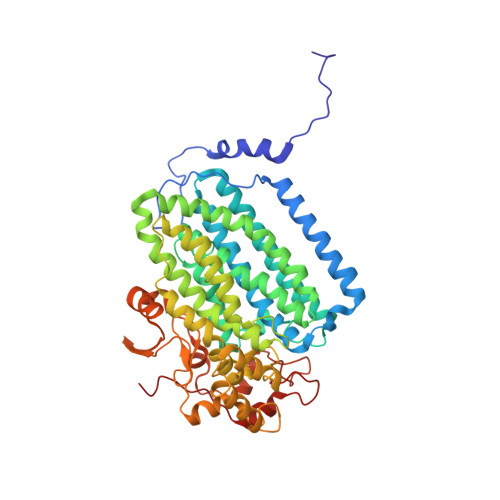
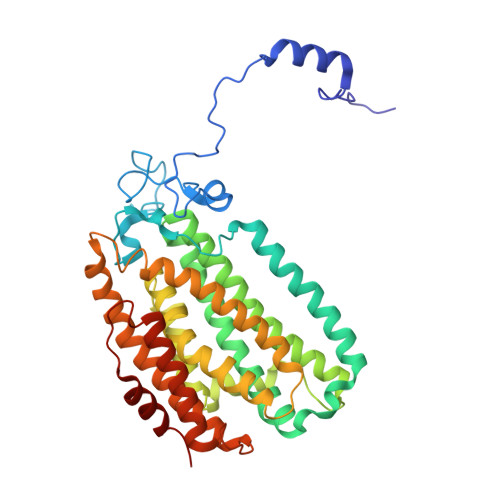
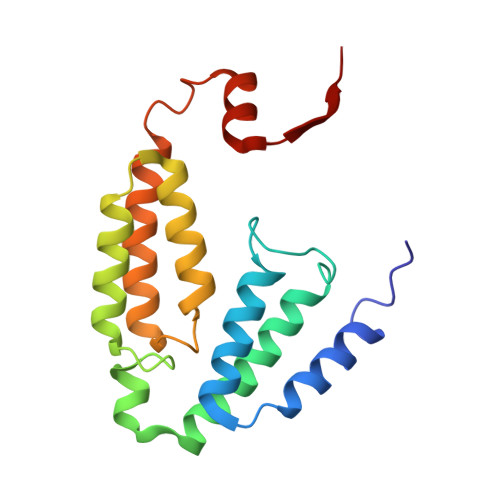
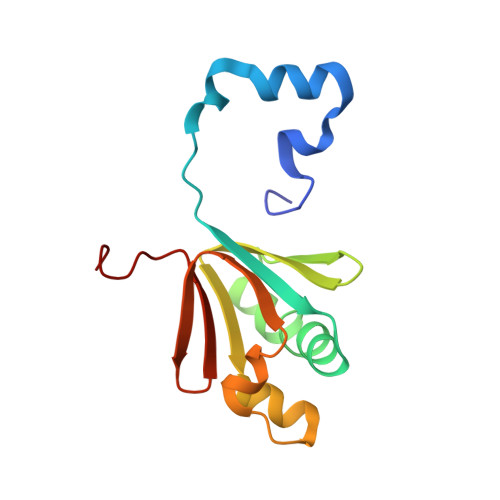
X-ray Crystal Structures of Methane Monooxygenase Hydroxylase Complexes with Variants of Its Regulatory Component: Correlations with Altered Reaction Cycle Dynamics.
Jones, J.C., Banerjee, R., Semonis, M.M., Shi, K., Aihara, H., Lipscomb, J.D.(2022) Biochemistry 61: 21-33
- PubMed: 34910460 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.1c00673
- Primary Citation Related Structures:
7S6Q, 7S6R, 7S6S, 7S6T, 7S7H - PubMed Abstract:
Full activity of soluble methane monooxygenase (sMMO) depends upon the formation of a 1:1 complex of the regulatory protein MMOB with each alpha subunit of the (αβγ) 2 hydroxylase, sMMOH. Previous studies have shown that mutations in the core region of MMOB and in the N- and C-termini cause dramatic changes in the rate constants for steps in the sMMOH reaction cycle. Here, X-ray crystal structures are reported for the sMMOH complex with two double variants within the core region of MMOB, DBL1 (N107G/S110A), and DBL2 (S109A/T111A), as well as two variants in the MMOB N-terminal region, H33A and H5A. DBL1 causes a 150-fold decrease in the formation rate constant of the reaction cycle intermediate P , whereas DBL2 accelerates the reaction of the dinuclear Fe(IV) intermediate Q with substrates larger than methane by three- to fourfold. H33A also greatly slows P formation, while H5A modestly slows both formation of Q and its reactions with substrates. Complexation with DBL1 or H33A alters the position of sMMOH residue R245, which is part of a conserved hydrogen-bonding network encompassing the active site diiron cluster where P is formed. Accordingly, electron paramagnetic resonance spectra of sMMOH:DBL1 and sMMOH:H33A complexes differ markedly from that of sMMOH:MMOB, showing an altered electronic environment. In the sMMOH:DBL2 complex, the position of M247 in sMMOH is altered such that it enlarges a molecular tunnel associated with substrate entry into the active site. The H5A variant causes only subtle structural changes despite its kinetic effects, emphasizing the precise alignment of sMMOH and MMOB required for efficient catalysis.
- Department of Biochemistry, Molecular Biology, and Biophysics, University of Minnesota, Minneapolis, Minnesota 55455, United States.
Organizational Affiliation: