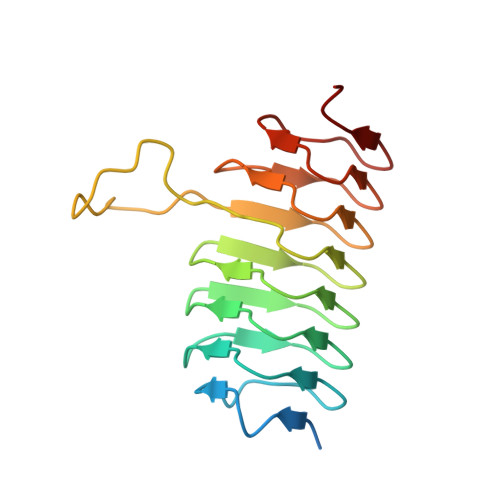
Biochemical investigation of an N-acetyltransferase from Helicobacter pullorum.
Griffiths, W.A., Spencer, K.D., Thoden, J.B., Holden, H.M.(2021) Protein Sci 30: 2418-2432
- PubMed: 34651380 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.4207
- Primary Citation Related Structures:
7S3U, 7S3W, 7S41, 7S42, 7S43, 7S44, 7S45 - PubMed Abstract:
N-acetylated sugars are often found, for example, on the lipopolysaccharides of Gram-negative bacteria, on the S-layers of Gram-positive bacteria, and on the capsular polysaccharides. Key enzymes involved in their biosynthesis are the sugar N-acetyltransferases. Here, we describe a structural and functional analysis of one such enzyme from Helicobacter pullorum, an emerging pathogen that may be associated with gastroenteritis and gallbladder and liver diseases. For this analysis, the gene BA919-RS02330 putatively encoding an N-acetyltransferase was cloned, and the corresponding protein was expressed and purified. A kinetic analysis demonstrated that the enzyme utilizes dTDP-3-amino-3,6-dideoxy-d-glucose as a substrate as well as dTDP-3-amino-3,6-dideoxy-d-galactose, albeit at a reduced rate. In addition to this kinetic analysis, a similar enzyme from Helicobacter bilis was cloned and expressed, and its kinetic parameters were determined. Seven X-ray crystallographic structures of various complexes of the H. pullorum wild-type enzyme (or the C80T variant) were determined to resolutions of 1.7 Å or higher. The overall molecular architecture of the H. pullorum N-acetyltransferase places it into the Class II left-handed-β-helix superfamily (LβH). Taken together, the data presented herein suggest that 3-acetamido-3,6-dideoxy-d-glucose (or the galactose derivative) is found on either the H. pullorum O-antigen or in another of its complex glycoconjugates. A BLAST search suggests that more than 50 non-pylori Helicobacter spp. have genes encoding N-acetyltransferases. Given that there is little information concerning the complex glycans in non-pylori Helicobacter spp. and considering their zoonotic potential, our results provide new biochemical insight into these pathogens.
- Department of Biochemistry, University of Wisconsin, Madison, Wisconsin, USA.
Organizational Affiliation: