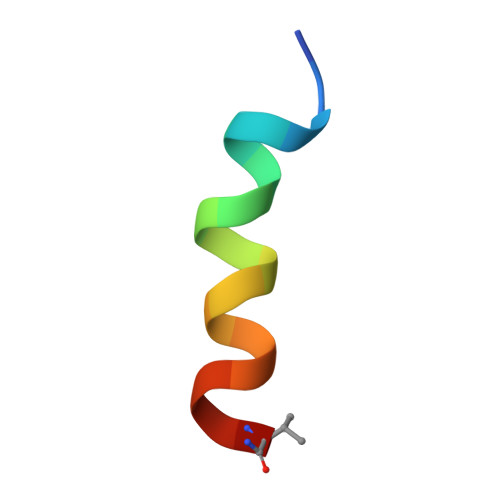
Structural insight into the mechanisms underlying the membrane activity of a family of antimicrobial Uperin 3 peptides
Prasad, A.K., Tiwari, C., Ray, S., Holden, S., Armstrong, D.A., Rosengren, K.J., Rodger, A., Martin, L.L., Panwar, A.S.To be published.