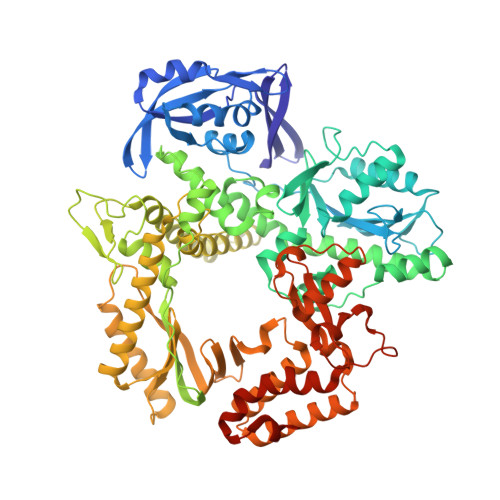
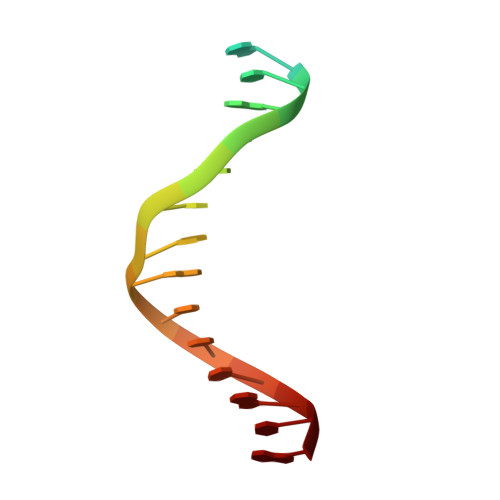
Synthesis and Polymerase Recognition of Threose Nucleic Acid Triphosphates Equipped with Diverse Chemical Functionalities.
Li, Q., Maola, V.A., Chim, N., Hussain, J., Lozoya-Colinas, A., Chaput, J.C.(2021) J Am Chem Soc 143: 17761-17768
- PubMed: 34637287 Search on PubMed
- DOI: https://doi.org/10.1021/jacs.1c08649
- Primary Citation Related Structures:
7RSU - PubMed Abstract:
Expanding the chemical space of evolvable non-natural genetic polymers (XNAs) to include functional groups that enhance protein target binding affinity offers a promising route to therapeutic aptamers with high biological stability. Here we describe the chemical synthesis and polymerase recognition of 10 chemically diverse functional groups introduced at the C-5 position of α-l-threofuranosyl uridine nucleoside triphosphate (tUTP). We show that the set of tUTP substrates is universally recognized by the laboratory-evolved polymerase Kod-RSGA. Insights into the mechanism of TNA synthesis were obtained from a high-resolution X-ray crystal structure of the postcatalytic complex bound to the primer-template duplex. A structural analysis reveals a large cavity in the enzyme active site that can accommodate the side chain of C-5-modified tUTP substrates. Our findings expand the chemical space of evolvable nucleic acid systems by providing a synthetic route to artificial genetic polymers that are uniformly modified with diversity-enhancing functional groups.