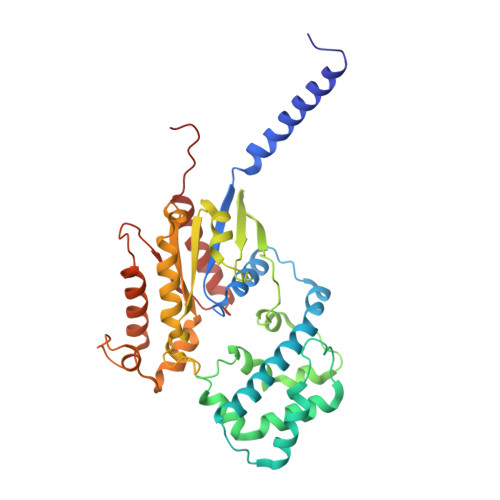
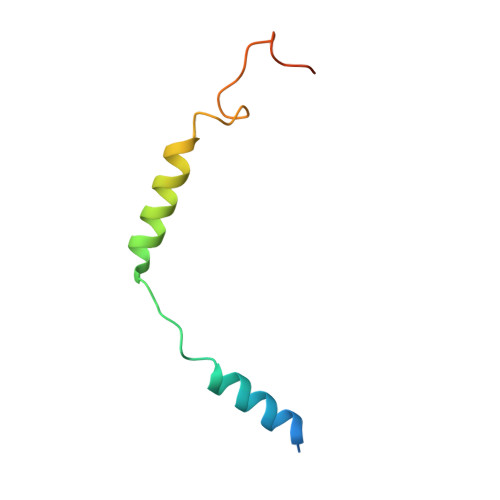
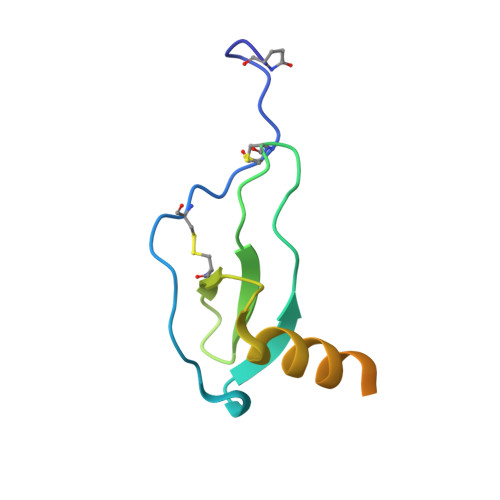
Atypical structural snapshots of human cytomegalovirus GPCR interactions with host G proteins
Tsutsumi, N., Maeda, S., Qu, Q., Voegele, M., Jude, K.M., Suomivuori, C.M., Panova, O., Waghray, D., Kato, H.E., Velasco, A., Dror, R.O., Skiniotis, G., Kobilka, B.K., Garcia, K.C.(2022) Sci Adv 8: eabl5442