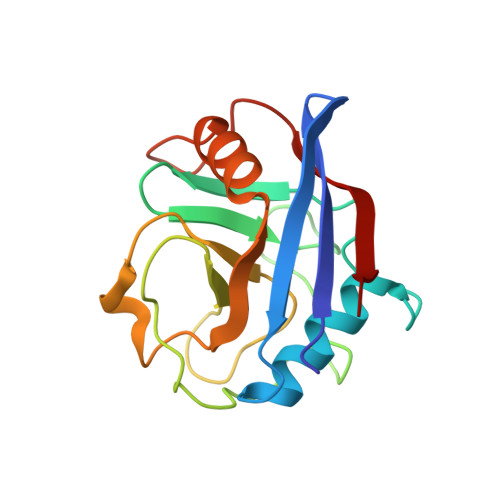
Structure-based design of pyrazole derivatives targeting the human Cyclophilin D binding site
Silva, D.O., Freitas, M.C., Malta, C.F., Martins, M.T., Sousa, P.M., Matias, P.M., Schwarz, D., Ventura, M.R., Gradler, U., Bandeiras, T.M.(2026) Int J Biol Macromol : 151496