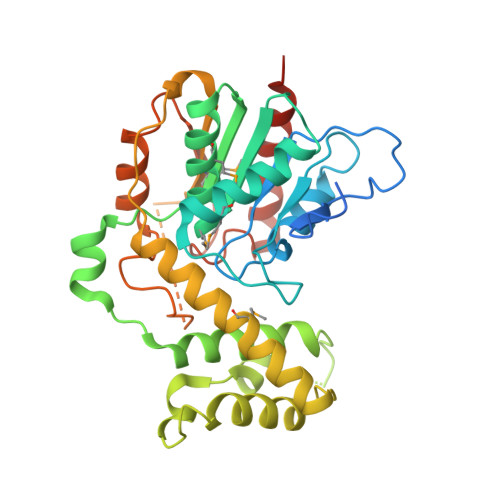
Structure of a Promiscuous Thioesterase Domain Responsible for Branching Acylation in Polyketide Biosynthesis.
Fraley, A.E., Dieterich, C.L., Mabesoone, M.F.J., Minas, H.A., Meoded, R.A., Hemmerling, F., Piel, J.(2022) Angew Chem Int Ed Engl 61: e202206385-e202206385
- PubMed: 35903999 Search on PubMed
- DOI: https://doi.org/10.1002/anie.202206385
- Primary Citation Related Structures:
7R0X - PubMed Abstract:
Thioesterases (TEs) are fundamentally important enzymes present in all bacteria and eukaryotes, where they have conserved functions in fatty acid biosynthesis and secondary metabolism. This work provides the first structural insights into a functionally distinct group of TEs that perform diverse acylations in polyketide and peptide biosynthesis (TE B s). Structural analysis of the oocydin (OocS) TE B domain facilitated identification and engineering of the active site to modulate acyl-group acceptance. In this way, we achieved higher reactivity using a structure-based approach, building a foundation for biocatalytic development of TE B -mediated O-acylation, a modification known to improve the bioactivity of oocydin-type polyketides. Lastly, the promiscuity of the OocS TE B motivated us to investigate, and ultimately provide evidence for, the production of longer chain branched oocydins in the native host Serratia plymuthica 4Rx13. This work frames the OocS TE B and homologs as invaluable synthetic biology tools for polyketide drug development.
- Department of Biology, Institute of Microbiology, Eidgenössische Technische Hochschule (ETH) Zurich, Vladimir-Prelog-Weg 4, 8093, Zurich, Switzerland.
Organizational Affiliation: