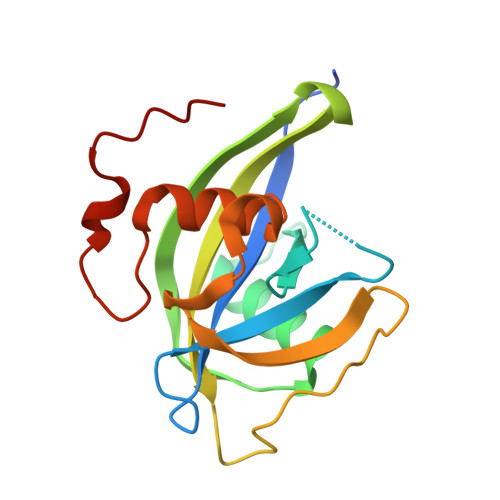
Kinetic and structural characterization of NUDT15 and NUDT18 as catalysts of isoprene pyrophosphate hydrolysis.
Scaletti, E.R., Unterlass, J.E., Almlof, I., Koolmeister, T., Vallin, K.S., Kapsitidou, D., Tsuber, V., Helleday, T., Stenmark, P., Jemth, A.S.(2024) FEBS J
- PubMed: 38944687 Search on PubMed
- DOI: https://doi.org/10.1111/febs.17202
- Primary Citation Related Structures:
7R0D - PubMed Abstract:
Isoprene pyrophosphates play a crucial role in the synthesis of a diverse array of essential nonsterol and sterol biomolecules and serve as substrates for posttranslational isoprenylation of proteins, enabling specific anchoring to cellular membranes. Hydrolysis of isoprene pyrophosphates would be a means to modulate their levels, downstream products, and protein isoprenylation. While NUDIX hydrolases from plants have been described to catalyze the hydrolysis of isoprene pyrophosphates, homologous enzymes with this function in animals have not yet been reported. In this study, we screened an extensive panel of human NUDIX hydrolases for activity in hydrolyzing isoprene pyrophosphates. We found that human nucleotide triphosphate diphosphatase NUDT15 and 8-oxo-dGDP phosphatase NUDT18 efficiently catalyze the hydrolysis of several physiologically relevant isoprene pyrophosphates. Notably, we demonstrate that geranyl pyrophosphate is an excellent substrate for NUDT18, with a catalytic efficiency of 2.1 × 10 5 m -1 ·s -1 , thus making it the best substrate identified for NUDT18 to date. Similarly, geranyl pyrophosphate proved to be the best isoprene pyrophosphate substrate for NUDT15, with a catalytic efficiency of 4.0 × 10 4 M -1 ·s -1 . LC-MS analysis of NUDT15 and NUDT18 catalyzed isoprene pyrophosphate hydrolysis revealed the generation of the corresponding monophosphates and inorganic phosphate. Furthermore, we solved the crystal structure of NUDT15 in complex with the hydrolysis product geranyl phosphate at a resolution of 1.70 Å. This structure revealed that the active site nicely accommodates the hydrophobic isoprenoid moiety and helped identify key binding residues. Our findings imply that isoprene pyrophosphates are endogenous substrates of NUDT15 and NUDT18, suggesting they are involved in animal isoprene pyrophosphate metabolism.
- Department of Biochemistry and Biophysics, Stockholm University, Sweden.
Organizational Affiliation: