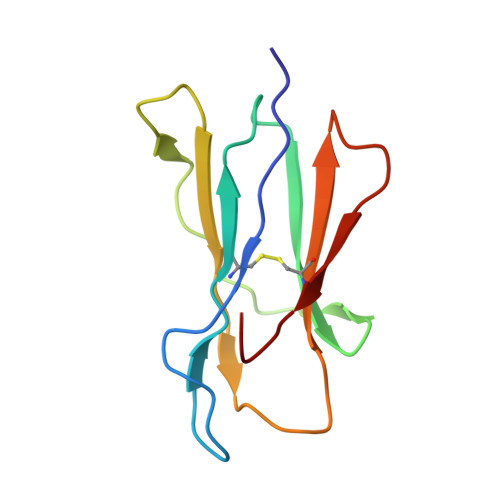
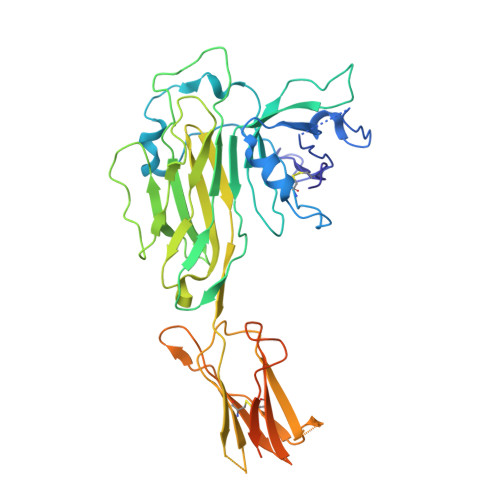
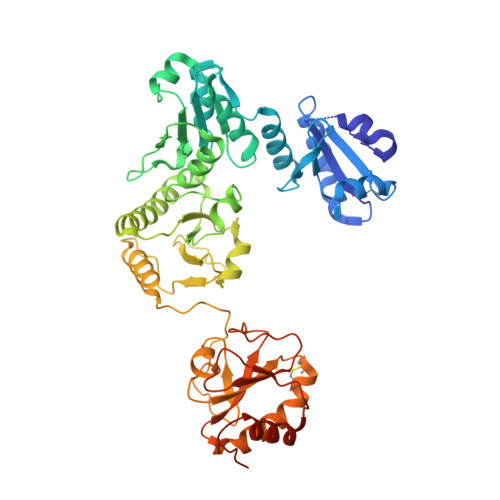
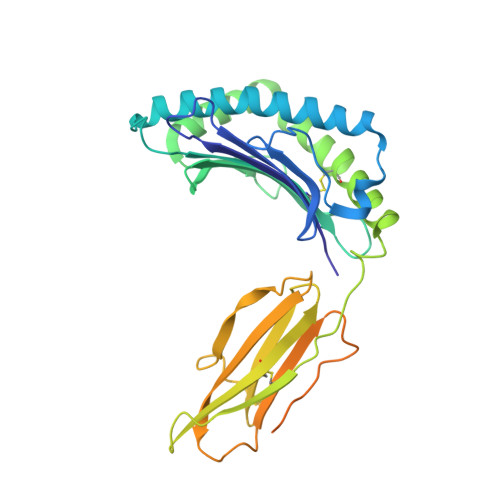
Molecular basis of MHC I quality control in the peptide loading complex.
Domnick, A., Winter, C., Susac, L., Hennecke, L., Hensen, M., Zitzmann, N., Trowitzsch, S., Thomas, C., Tampe, R.(2022) Nat Commun 13: 4701-4701
- PubMed: 35948544 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-32384-z
- Primary Citation Related Structures:
7QPD - PubMed Abstract:
Major histocompatibility complex class I (MHC I) molecules are central to adaptive immunity. Their assembly, epitope selection, and antigen presentation are controlled by the MHC I glycan through a sophisticated network of chaperones and modifying enzymes. However, the mechanistic integration of the corresponding processes remains poorly understood. Here, we determine the multi-chaperone-client interaction network of the peptide loading complex (PLC) and report the PLC editing module structure by cryogenic electron microscopy at 3.7 Å resolution. Combined with epitope-proofreading studies of the PLC in near-native lipid environment, these data show that peptide-receptive MHC I molecules are stabilized by multivalent chaperone interactions including the calreticulin-engulfed mono-glucosylated MHC I glycan, which only becomes accessible for processing by α-glucosidase II upon loading of optimal epitopes. Our work reveals allosteric coupling between peptide-MHC I assembly and glycan processing. This inter-process communication defines the onset of an adaptive immune response and provides a prototypical example of the tightly coordinated events in endoplasmic reticulum quality control.
- Institute of Biochemistry, Biocenter, Goethe University Frankfurt, Max-von-Laue-Str. 9, 60438, Frankfurt am Main, Germany.
Organizational Affiliation: