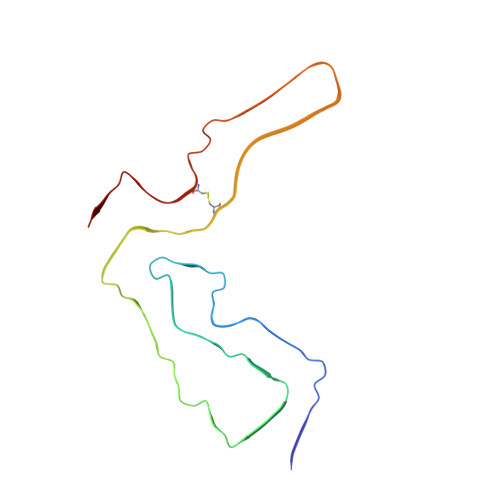
2.7 angstrom cryo-EM structure of ex vivo RML prion fibrils.
Manka, S.W., Zhang, W., Wenborn, A., Betts, J., Joiner, S., Saibil, H.R., Collinge, J., Wadsworth, J.D.F.(2022) Nat Commun 13: 4004-4004
- PubMed: 35831275 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-30457-7
- Primary Citation Related Structures:
7QIG - PubMed Abstract:
Mammalian prions propagate as distinct strains and are composed of multichain assemblies of misfolded host-encoded prion protein (PrP). Here, we present a near-atomic resolution cryo-EM structure of PrP fibrils present in highly infectious prion rod preparations isolated from the brains of RML prion-infected mice. We found that prion rods comprise single-protofilament helical amyloid fibrils that coexist with twisted pairs of the same protofilaments. Each rung of the protofilament is formed by a single PrP monomer with the ordered core comprising PrP residues 94-225, which folds to create two asymmetric lobes with the N-linked glycans and the glycosylphosphatidylinositol anchor projecting from the C-terminal lobe. The overall architecture is comparable to that of recently reported PrP fibrils isolated from the brain of hamsters infected with the 263K prion strain. However, there are marked conformational variations that could result from differences in PrP sequence and/or represent distinguishing features of the distinct prion strains.
- MRC Prion Unit at UCL, Institute of Prion Diseases, University College London, 33 Cleveland Street, London, W1W 7FF, UK.
Organizational Affiliation: