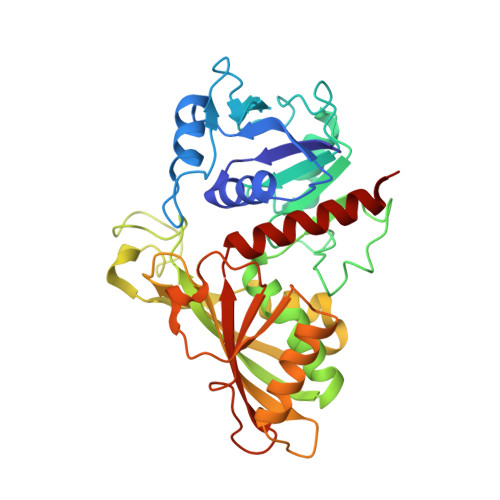
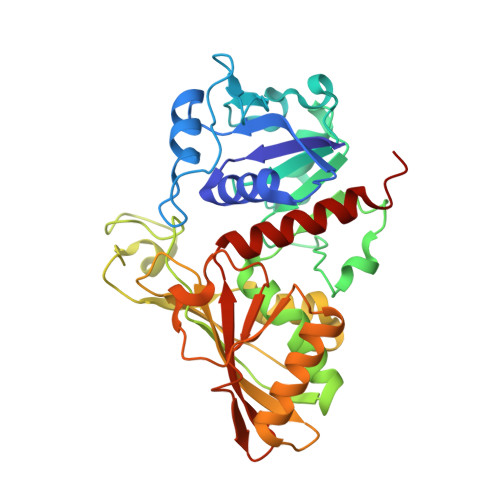
Unravelling the regulation pathway of photosynthetic AB-GAPDH.
Marotta, R., Del Giudice, A., Gurrieri, L., Fanti, S., Swuec, P., Galantini, L., Falini, G., Trost, P., Fermani, S., Sparla, F.(2022) Acta Crystallogr D Struct Biol 78: 1399-1411
- PubMed: 36322422 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798322010014
- Primary Citation Related Structures:
7Q53, 7Q54, 7Q55, 7Q56, 7Q57 - PubMed Abstract:
Oxygenic phototrophs perform carbon fixation through the Calvin-Benson cycle. Different mechanisms adjust the cycle and the light-harvesting reactions to rapid environmental changes. Photosynthetic glyceraldehyde 3-phosphate dehydrogenase (GAPDH) is a key enzyme in the cycle. In land plants, different photosynthetic GAPDHs exist: the most abundant isoform is formed by A 2 B 2 heterotetramers and the least abundant by A 4 homotetramers. Regardless of the subunit composition, GAPDH is the major consumer of photosynthetic NADPH and its activity is strictly regulated. While A 4 -GAPDH is regulated by CP12, AB-GAPDH is autonomously regulated through the C-terminal extension (CTE) of its B subunits. Reversible inhibition of AB-GAPDH occurs via the oxidation of a cysteine pair located in the CTE and the substitution of NADP(H) with NAD(H) in the cofactor-binding site. These combined conditions lead to a change in the oligomerization state and enzyme inhibition. SEC-SAXS and single-particle cryo-EM analysis were applied to reveal the structural basis of this regulatory mechanism. Both approaches revealed that spinach (A 2 B 2 ) n -GAPDH oligomers with n = 1, 2, 4 and 5 co-exist in a dynamic system. B subunits mediate the contacts between adjacent tetramers in A 4 B 4 and A 8 B 8 oligomers. The CTE of each B subunit penetrates into the active site of a B subunit of the adjacent tetramer, which in turn moves its CTE in the opposite direction, effectively preventing the binding of the substrate 1,3-bisphosphoglycerate in the B subunits. The whole mechanism is made possible, and eventually controlled, by pyridine nucleotides. In fact, NAD(H), by removing NADP(H) from A subunits, allows the entrance of the CTE into the active site of the B subunit, hence stabilizing inhibited oligomers.
- Electron Microscopy Facility (EMF), Italian Institute of Technology (IIT), 16163 Genova, Italy.
Organizational Affiliation: