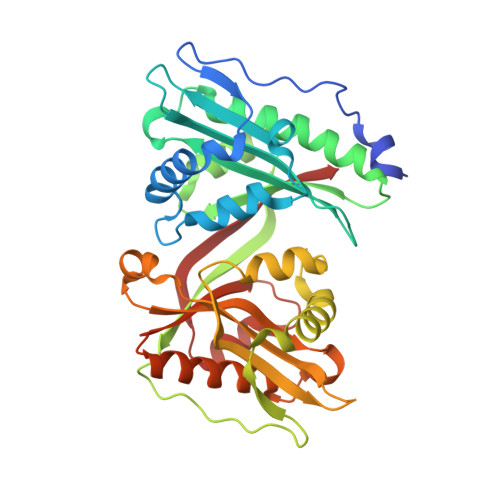
Biochemical, structural, and functional studies reveal that MAB_4324c from Mycobacterium abscessus is an active tandem repeat N-acetyltransferase.
Alsarraf, H.M.A.B., Ung, K.L., Johansen, M.D., Dimon, J., Olieric, V., Kremer, L., Blaise, M.(2022) FEBS Lett 596: 1516-1532
- PubMed: 35470425 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.14360
- Primary Citation Related Structures:
7Q3A - PubMed Abstract:
Mycobacterium abscessus is a pathogenic non-tuberculous mycobacterium that possesses an intrinsic drug resistance profile. Several N-acetyltransferases mediate drug resistance and/or participate in M. abscessus virulence. Mining the M. abscessus genome has revealed genes encoding additional N-acetyltransferases whose functions remain uncharacterized, among them MAB_4324c. Here, we showed that the purified MAB_4324c protein is a N-acetyltransferase able to acetylate small polyamine substrates. The crystal structure of MAB_4324c was solved at high resolution in complex with its cofactor, revealing the presence of two GCN5-related N-acetyltransferase domains and a cryptic binding site for NADPH. Genetic studies demonstrate that MAB_4324c is not essential for in vitro growth of M. abscessus; however, overexpression of the protein enhanced the uptake and survival of M. abscessus in THP-1 macrophages.
- IRIM, CNRS, Université de Montpellier, France.
Organizational Affiliation: