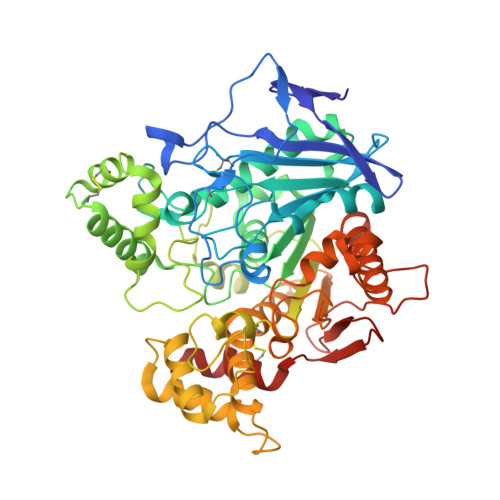
Discovery of new, highly potent and selective inhibitors of BuChE - design, synthesis, in vitro and in vivo evaluation and crystallography studies.
Panek, D., Pasieka, A., Latacz, G., Zareba, P., Szczech, M., Godyn, J., Chantegreil, F., Nachon, F., Brazzolotto, X., Skrzypczak-Wiercioch, A., Walczak, M., Smolik, M., Salat, K., Hofner, G., Wanner, K., Wieckowska, A., Malawska, B.(2023) Eur J Med Chem 249: 115135-115135
- PubMed: 36696766 Search on PubMed
- DOI: https://doi.org/10.1016/j.ejmech.2023.115135
- Primary Citation Related Structures:
7Q1M, 7Q1N, 7Q1O, 7Q1P - PubMed Abstract:
The symptomatic and disease-modifying effects of butyrylcholinesterase (BuChE) inhibitors provide an encouraging premise for researching effective treatments for Alzheimer's disease. Here, we examined a series of compounds with a new chemical scaffold based on 3-(cyclohexylmethyl)amino-2-hydroxypropyl, and we identified a highly selective hBuChE inhibitor (29). Based on extensive in vitro and in vivo evaluations of the compound and its enantiomers, (R)-29 was identified as a promising candidate for further development. Compound (R)-29 is a potent hBuChE inhibitor (IC 50 = 40 nM) with selectivity over AChE and relevant off-targets, including H 1 , M 1 , α 1A and β 1 receptors. The compound displays high metabolic stability on human liver microsomes (90% of the parent compound after 2 h of incubation), and its safety was confirmed through examining the cytotoxicity on the HepG2 cell line (LC 50 = 2.85 μM) and hERG inhibition (less than 50% at 10 μM). While (rac)-29 lacked an effect in vivo and showed limited penetration to the CNS in pharmacokinetics studies, compound (R)-29 exhibited a procognitive effect at 15 mg/kg in the passive avoidance task in scopolamine-treated mice.
- Department of Physicochemical Drug Analysis, Faculty of Pharmacy, Jagiellonian University Medical College, Medyczna St. 9, 30-688, Kraków, Poland. Electronic address: dawid.panek@uj.edu.pl.
Organizational Affiliation: