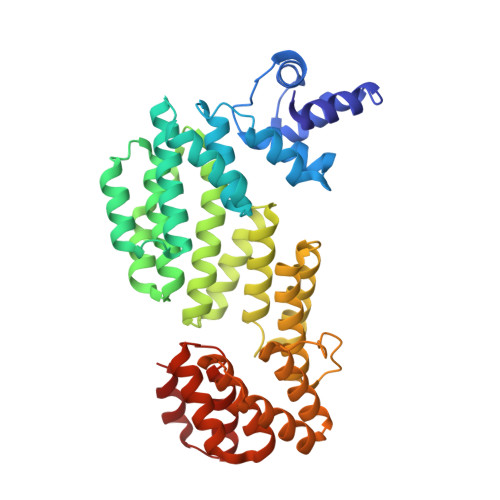
Insights into the mechanism of action of the arbitrium communication system in SPbeta phages.
Gallego Del Sol, F., Quiles-Puchalt, N., Brady, A., Penades, J.R., Marina, A.(2022) Nat Commun 13: 3627-3627
- PubMed: 35750663 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-31144-3
- Primary Citation Related Structures:
6S7I, 6S7L, 7Q0N - PubMed Abstract:
The arbitrium system is employed by phages of the SPbeta family to communicate with their progeny during infection to decide either to follow the lytic or the lysogenic cycle. The system is controlled by a peptide, AimP, that binds to the regulator AimR, inhibiting its DNA-binding activity and expression of aimX. Although the structure of AimR has been elucidated for phages SPβ and phi3T, there is still controversy regarding the molecular mechanism of AimR function, with two different proposed models for SPβ. In this study, we deepen our understanding of the system by solving the structure of an additional AimR that shows chimerical characteristics with the SPβ receptor. The crystal structures of this AimR (apo, AimP-bound and DNA-bound) together with in vitro and in vivo analyses confirm a mechanism of action by AimP-induced conformational restriction, shedding light on peptide specificity and cross regulation with relevant biological implications.
- Instituto de Biomedicina de Valencia (IBV), CSIC and CIBER de Enfermedades Raras (CIBERER), Valencia, Spain.
Organizational Affiliation: