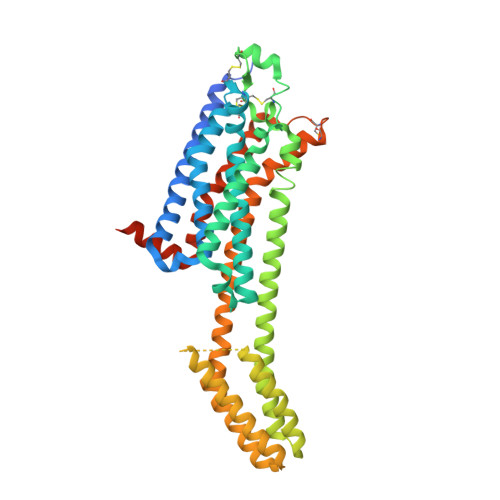
Single Stabilizing Point Mutation Enables High-Resolution Co-Crystal Structures of the Adenosine A 2A Receptor with Preladenant Conjugates.
Claff, T., Klapschinski, T.A., Tiruttani Subhramanyam, U.K., Vaassen, V.J., Schlegel, J.G., Vielmuth, C., Voss, J.H., Labahn, J., Muller, C.E.(2022) Angew Chem Int Ed Engl 61: e202115545-e202115545
- PubMed: 35174942 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/anie.202115545
- Primary Citation Related Structures:
7PX4, 7PYR - PubMed Abstract:
The G protein-coupled adenosine A 2A receptor (A 2A AR) is an important new (potential) drug target in immuno-oncology, and for neurodegenerative diseases. Preladenant and its derivatives belong to the most potent A 2A AR antagonists displaying exceptional selectivity. While crystal structures of the human A 2A AR have been solved, mostly using the A 2A -StaR2 protein that bears 9 point mutations, co-crystallization with Preladenant derivatives has so far been elusive. We developed a new A 2A AR construct harboring a single point mutation (S91 3.39 K) which renders it extremely thermostable. This allowed the co-crystallization of two novel Preladenant derivatives, the polyethylene glycol-conjugated (PEGylated) PSB-2113, and the fluorophore-labeled PSB-2115. The obtained crystal structures (2.25 Å and 2.6 Å resolution) provide explanations for the high potency and selectivity of Preladenant derivatives. They represent the first crystal structures of a GPCR in complex with PEG- and fluorophore-conjugated ligands. The applied strategy is predicted to be applicable to further class A GPCRs.
- Pharmaceutical Institute, Pharmaceutical & Medicinal Chemistry, University of Bonn, An der Immenburg 4, 53121, Bonn, Germany.
Organizational Affiliation: