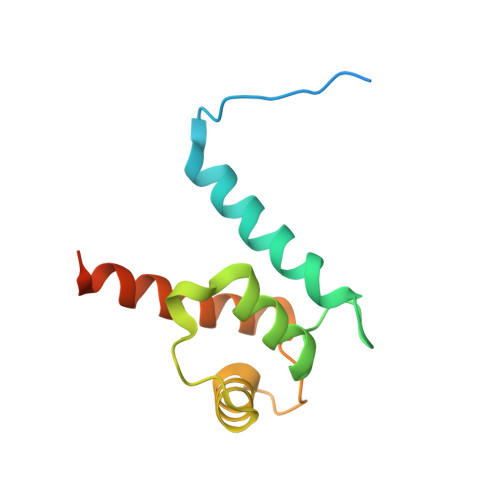
Crystal structure of S100A4 labeled with NU074381b.
Giroud, C., Szommer, T., Coxon, C., Monteiro, O., Christott, T., Bennett, J., Aitmakhanova, K., Raux, B., Newman, J., Elkins, J., Krojer, T., Koekemoer, L., Von Delft, F., Bountr, C., Brennan, P., Fedorov, O.To be published.