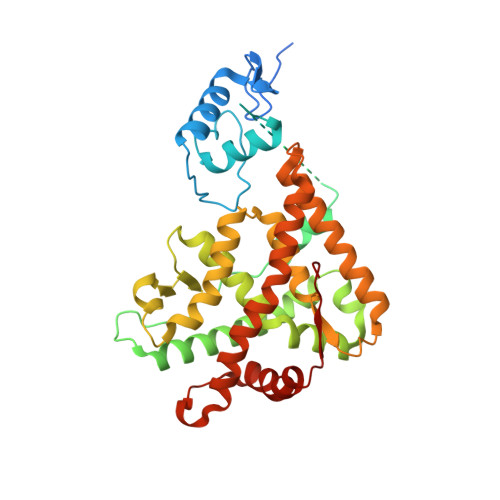
Quaternary glucocorticoid receptor structure highlights allosteric interdomain communication.
Postel, S., Wissler, L., Johansson, C.A., Gunnarsson, A., Gordon, E., Collins, B., Castaldo, M., Kohler, C., Oling, D., Johansson, P., Froderberg Roth, L., Beinsteiner, B., Dainty, I., Delaney, S., Klaholz, B.P., Billas, I.M.L., Edman, K.(2023) Nat Struct Mol Biol 30: 286-295
- PubMed: 36747092 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-022-00914-4
- Primary Citation Related Structures:
7PRV, 7PRW, 7PRX - PubMed Abstract:
The glucocorticoid receptor (GR) is a ligand-activated transcription factor that binds DNA and assembles co-regulator complexes to regulate gene transcription. GR agonists are widely prescribed to people with inflammatory and autoimmune diseases. Here we present high-resolution, multidomain structures of GR in complex with ligand, DNA and co-regulator peptide. The structures reveal how the receptor forms an asymmetric dimer on the DNA and provide a detailed view of the domain interactions within and across the two monomers. Hydrogen-deuterium exchange and DNA-binding experiments demonstrate that ligand-dependent structural changes are communicated across the different domains in the full-length receptor. This study demonstrates how GR forms a distinct architecture on DNA and how signal transmission can be modulated by the ligand pharmacophore, provides a platform to build a new level of understanding of how receptor modifications can drive disease progression and offers key insight for future drug design.
- Mechanistic & Structural Biology, Discovery Sciences, R&D, AstraZeneca, Gothenburg, Sweden.
Organizational Affiliation: