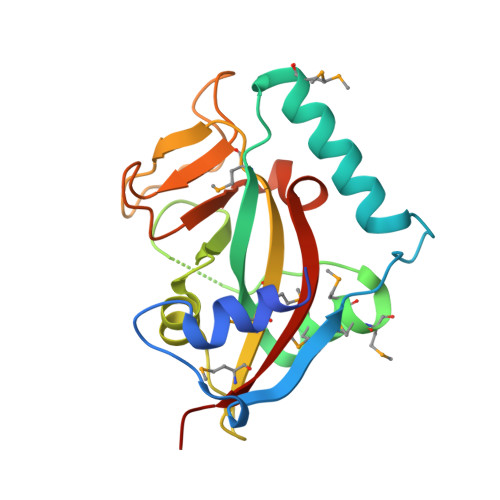
The superior salinity tolerance of bread wheat cultivar Shanrong No. 3 is unlikely to be caused by elevated Ta-sro1 poly-(ADP-ribose) polymerase activity.
Vogt, S., Feijs, K., Hosch, S., De Masi, R., Lintermann, R., Loll, B., Wirthmueller, L.(2022) Plant Cell 34: 4130-4137
- PubMed: 35980152 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/plcell/koac261
- Primary Citation Related Structures:
7PLQ - PubMed Abstract:
Structural and biochemical analyses demonstrate that the elevated salinity tolerance of bread wheat cultivar Shanrong No. 3 is unlikely to be caused by elevated Ta-sro1 poly(ADP-ribose) polymerase activity.
- Department Biochemistry of Plant Interactions, Leibniz Institute of Plant Biochemistry, Halle (Saale), 06120, Germany.
Organizational Affiliation: