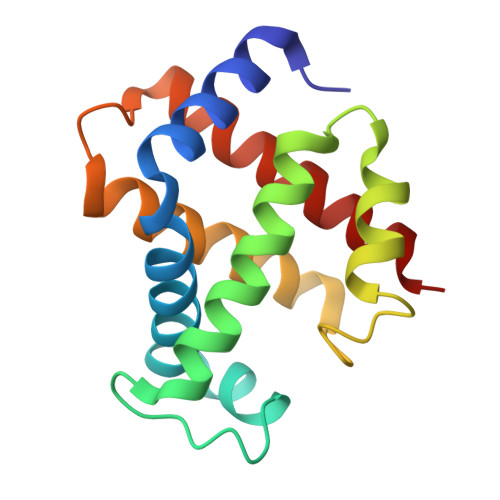
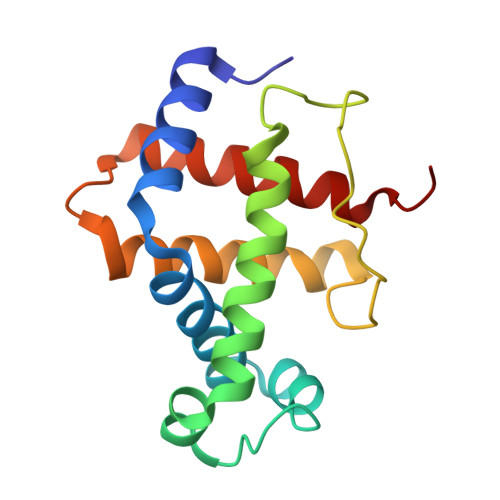
Cryo-EM structures of staphylococcal IsdB bound to human hemoglobin reveal the process of heme extraction.
De Bei, O., Marchetti, M., Ronda, L., Gianquinto, E., Lazzarato, L., Chirgadze, D.Y., Hardwick, S.W., Cooper, L.R., Spyrakis, F., Luisi, B.F., Campanini, B., Bettati, S.(2022) Proc Natl Acad Sci U S A 119: e2116708119-e2116708119
- PubMed: 35357971 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2116708119
- Primary Citation Related Structures:
7PCF, 7PCH, 7PCQ - PubMed Abstract:
Iron surface determinant B (IsdB) is a hemoglobin (Hb) receptor essential for hemic iron acquisition by Staphylococcus aureus. Heme transfer to IsdB is possible from oxidized Hb (metHb), but inefficient from Hb either bound to oxygen (oxyHb) or bound to carbon monoxide (HbCO), and encompasses a sequence of structural events that are currently poorly understood. By single-particle cryo-electron microscopy, we determined the structure of two IsdB:Hb complexes, representing key species along the heme extraction pathway. The IsdB:HbCO structure, at 2.9-Å resolution, provides a snapshot of the preextraction complex. In this early stage of IsdB:Hb interaction, the hemophore binds to the β-subunits of the Hb tetramer, exploiting a folding-upon-binding mechanism that is likely triggered by a cis/trans isomerization of Pro173. Binding of IsdB to α-subunits occurs upon dissociation of the Hb tetramer into α/β dimers. The structure of the IsdB:metHb complex reveals the final step of the extraction process, where heme transfer to IsdB is completed. The stability of the complex, both before and after heme transfer from Hb to IsdB, is influenced by isomerization of Pro173. These results greatly enhance current understanding of structural and dynamic aspects of the heme extraction mechanism by IsdB and provide insight into the interactions that stabilize the complex before the heme transfer event. This information will support future efforts to identify inhibitors of heme acquisition by S. aureus by interfering with IsdB:Hb complex formation.
- Interdepartmental Center Biopharmanet-TEC, University of Parma, Parma 43124, Italy.
Organizational Affiliation: