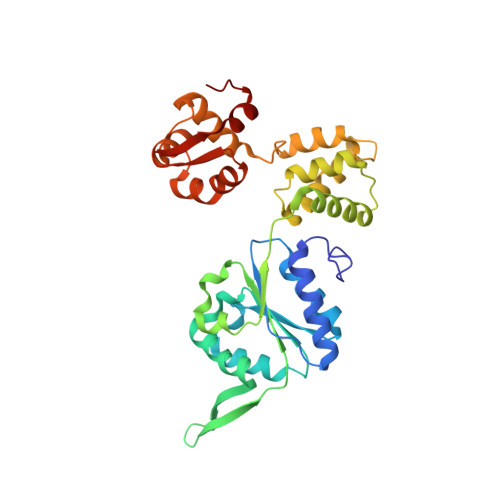
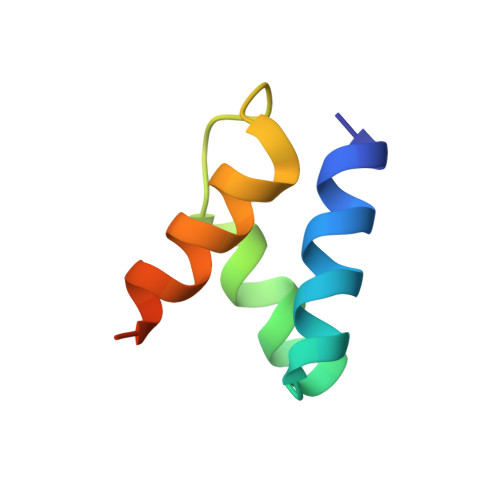
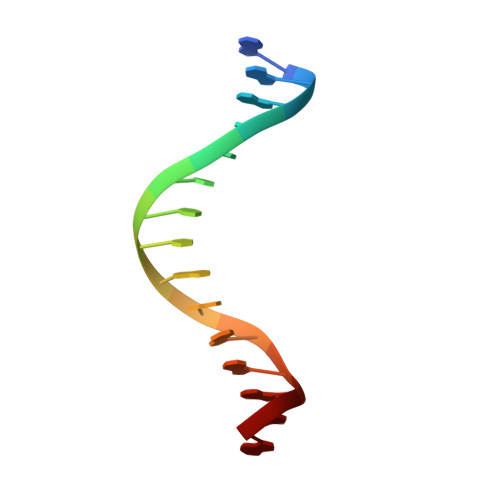
Mechanism of AAA+ ATPase-mediated RuvAB-Holliday junction branch migration.
Wald, J., Fahrenkamp, D., Goessweiner-Mohr, N., Lugmayr, W., Ciccarelli, L., Vesper, O., Marlovits, T.C.(2022) Nature 609: 630-639
- PubMed: 36002576 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-022-05121-1
- Primary Citation Related Structures:
7PBL, 7PBM, 7PBN, 7PBO, 7PBP, 7PBQ, 7PBR, 7PBS, 7PBT, 7PBU - PubMed Abstract:
The Holliday junction is a key intermediate formed during DNA recombination across all kingdoms of life 1 . In bacteria, the Holliday junction is processed by two homo-hexameric AAA+ ATPase RuvB motors, which assemble together with the RuvA-Holliday junction complex to energize the strand-exchange reaction 2 . Despite its importance for chromosome maintenance, the structure and mechanism by which this complex facilitates branch migration are unknown. Here, using time-resolved cryo-electron microscopy, we obtained structures of the ATP-hydrolysing RuvAB complex in seven distinct conformational states, captured during assembly and processing of a Holliday junction. Five structures together resolve the complete nucleotide cycle and reveal the spatiotemporal relationship between ATP hydrolysis, nucleotide exchange and context-specific conformational changes in RuvB. Coordinated motions in a converter formed by DNA-disengaged RuvB subunits stimulate hydrolysis and nucleotide exchange. Immobilization of the converter enables RuvB to convert the ATP-contained energy into a lever motion, which generates the pulling force driving the branch migration. We show that RuvB motors rotate together with the DNA substrate, which, together with a progressing nucleotide cycle, forms the mechanistic basis for DNA recombination by continuous branch migration. Together, our data decipher the molecular principles of homologous recombination by the RuvAB complex, elucidate discrete and sequential transition-state intermediates for chemo-mechanical coupling of hexameric AAA+ motors and provide a blueprint for the design of state-specific compounds targeting AAA+ motors.
- Institute of Structural and Systems Biology, University Medical Center Hamburg-Eppendorf, Hamburg, Germany. jiri.wald@cssb-hamburg.de.
Organizational Affiliation: