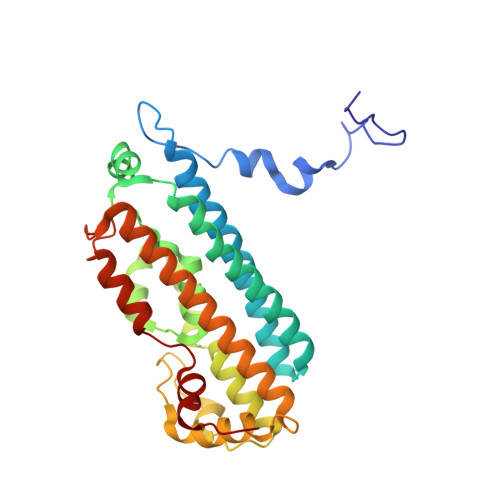
Binding of l-kynurenine to X. campestris tryptophan 2,3-dioxygenase.
Basran, J., Booth, E.S., Campbell, L.P., Thackray, S.J., Jesani, M.H., Clayden, J., Moody, P.C.E., Mowat, C.G., Kwon, H., Raven, E.L.(2021) J Inorg Biochem 225: 111604-111604
- PubMed: 34571402 Search on PubMed
- DOI: https://doi.org/10.1016/j.jinorgbio.2021.111604
- Primary Citation Related Structures:
7P46 - PubMed Abstract:
The kynurenine pathway is the major route of tryptophan metabolism. The first step of this pathway is catalysed by one of two heme-dependent dioxygenase enzymes - tryptophan 2,3-dioxygenase (TDO) and indoleamine 2,3-dioxygenase (IDO) - leading initially to the formation of N-formylkynurenine (NFK). In this paper, we present a crystal structure of a bacterial TDO from X. campestris in complex with l-kynurenine, the hydrolysed product of NFK. l-kynurenine is bound at the active site in a similar location to the substrate (l-Trp). Hydrogen bonding interactions with Arg117 and the heme 7-propionate anchor the l-kynurenine molecule into the pocket. A mechanism for the hydrolysis of NFK in the active site is presented.
- Department of Molecular and Cell Biology, Leicester Institute of Structural and Chemical Biology, University of Leicester, Leicester LE1 9HN, UK.
Organizational Affiliation: