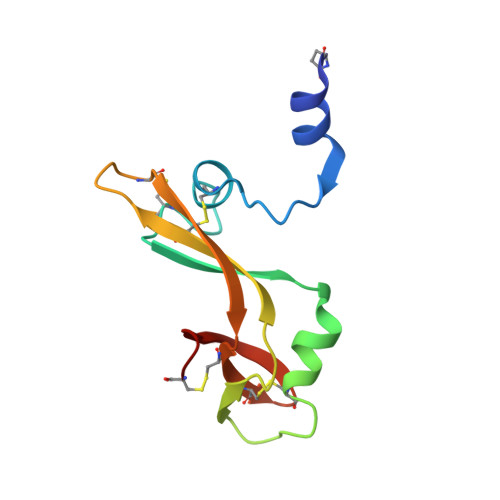
The crystal structure of the domain-swapped dimer of onconase highlights some catalytic and antitumor activity features of the enzyme.
Gotte, G., Campagnari, R., Loreto, D., Bettin, I., Calzetti, F., Menegazzi, M., Merlino, A.(2021) Int J Biol Macromol 191: 560-571
- PubMed: 34563576 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2021.09.095
- Primary Citation Related Structures:
7OR6, 7ORD - PubMed Abstract:
Onconase (ONC) is a monomeric amphibian "pancreatic-type" RNase endowed with remarkable anticancer activity. ONC spontaneously forms traces of a dimer (ONC-D) in solution, while larger amounts can be formed when ONC is lyophilized from mildly acidic solutions. Here, we report the crystal structure of ONC-D and analyze its catalytic and antitumor activities in comparison to ONC. ONC-D forms via the three-dimensional swapping of the N-terminal α-helix between two monomers, but it displays a significantly different quaternary structure from that previously modeled [Fagagnini A et al., 2017, Biochem J 474, 3767-81], and based on the crystal structure of the RNase A N-terminal swapped dimer. ONC-D presents a variable quaternary assembly deriving from a variable open interface, while it retains a catalytic activity that is similar to that of ONC. Notably, ONC-D displays antitumor activity against two human melanoma cell lines, although it exerts a slightly lower cytostatic effect than the monomer. The inhibition of melanoma cell proliferation by ONC or ONC-D is associated with the reduction of the expression of the anti-apoptotic B cell lymphoma 2 (Bcl2), as well as of the total expression and phosphorylation of the Signal Transducer and Activator of Transcription (STAT)-3. Phosphorylation is inhibited in both STAT3 Tyr705 and Ser727 key-residues, as well as in its upstream tyrosine-kinase Src. Consequently, both ONC species should exert their anti-cancer action by inhibiting the pro-tumor pleiotropic STAT3 effects deriving either by its phospho-tyrosine activation or by its non-canonical signaling pathways. Both ONC species, indeed, increase the portion of A375 cells undergoing apoptotic cell death. This study expands the variety of RNase domain-swapped dimeric structures, underlining the unpredictability of the open interface arrangement upon domain swapping. Structural data also offer valuable insights to analyze the differences in the measured ONC or ONC-D biological activities.
- Department of Neuroscience, Biomedicine and Movement Sciences, Biological Chemistry Section, University of Verona, Strada Le Grazie 8, 37134 Verona, Italy. Electronic address: giovanni.gotte@univr.it.
Organizational Affiliation: