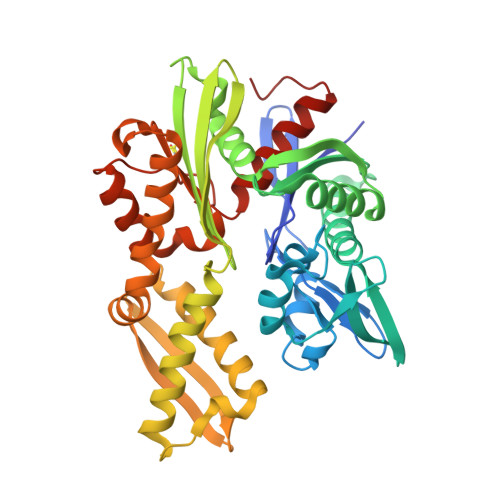
Structures of the Plasmodium falciparum heat-shock protein 70-x ATPase domain in complex with chemical fragments identify conserved and unique binding sites.
Mohamad, N., O'Donoghue, A., Kantsadi, A.L., Vakonakis, I.(2021) Acta Crystallogr F Struct Biol Commun 77: 262-268
- PubMed: 34341192 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X21007378
- Primary Citation Related Structures:
7OOE, 7OOG, 7P31 - PubMed Abstract:
Plasmodium falciparum invades erythrocytes and extensively modifies them in a manner that increases the virulence of this malaria parasite. A single heat-shock 70 kDa-type chaperone, PfHsp70-x, is among the parasite proteins exported to the host cell. PfHsp70-x assists in the formation of a key protein complex that underpins parasite virulence and supports parasite growth during febrile episodes. Previous work resolved the crystallographic structures of the PfHsp70-x ATPase and substrate-binding domains, and showed them to be highly similar to those of their human counterparts. Here, 233 chemical fragments were screened for binding to the PfHsp70-x ATPase domain, resulting in three crystallographic structures of this domain in complex with ligands. Two binding sites were identified, with most ligands binding proximal to the ATPase nucleotide-binding pocket. Although amino acids participating in direct ligand interactions are conserved between the parasite and human erythrocytic chaperones, one nonconserved residue is also present near the ligand. This work suggests that PfHsp70-x features binding sites that may be exploitable by small-molecule ligands towards the specific inhibition of the parasite chaperone.
- Department of Biochemistry, University of Oxford, South Parks Road, Oxford OX1 3QU, United Kingdom.
Organizational Affiliation: