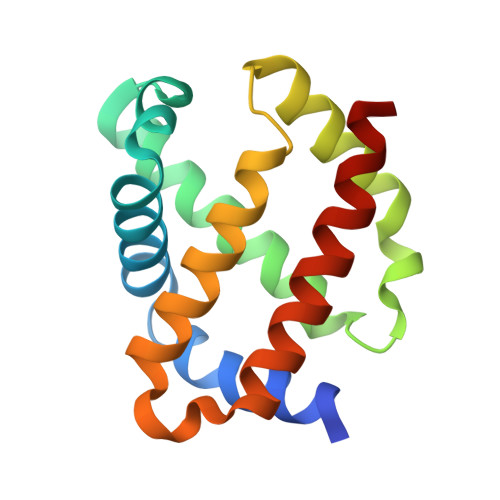
Probing the Role of Murine Neuroglobin CDloop-D-Helix Unit in CO Ligand Binding and Structural Dynamics.
Exertier, C., Sebastiani, F., Freda, I., Gugole, E., Cerutti, G., Parisi, G., Montemiglio, L.C., Becucci, M., Viappiani, C., Bruno, S., Savino, C., Zamparelli, C., Anselmi, M., Abbruzzetti, S., Smulevich, G., Vallone, B.(2022) ACS Chem Biol 17: 2099-2108
- PubMed: 35797699 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acschembio.2c00172
- Primary Citation Related Structures:
7OHD - PubMed Abstract:
We produced a neuroglobin variant, namely, Ngb CDless, with the excised CDloop- and D-helix, directly joining the C- and E-helices. The CDless variant retained bis-His hexacoordination, and we investigated the role of the CDloop-D-helix unit in controlling the CO binding and structural dynamics by an integrative approach based on X-ray crystallography, rapid mixing, laser flash photolysis, resonance Raman spectroscopy, and molecular dynamics simulations. Rapid mixing and laser flash photolysis showed that ligand affinity was unchanged with respect to the wild-type protein, albeit with increased on and off constants for rate-limiting heme iron hexacoordination by the distal His64. Accordingly, resonance Raman spectroscopy highlighted a more open distal pocket in the CO complex that, in agreement with MD simulations, likely involves His64 swinging inward and outward of the distal heme pocket. Ngb CDless displays a more rigid overall structure with respect to the wild type, abolishing the structural dynamics of the CDloop-D-helix hypothesized to mediate its signaling role, and it retains ligand binding control by distal His64. In conclusion, this mutant may represent a tool to investigate the involvement of CDloop-D-helix in neuroprotective signaling in a cellular or animal model.
- Dipartimento di Scienze Biochimiche "A. Rossi Fanelli", Sapienza, Università di Roma, Piazzale A. Moro 5, I-00185 Rome, Italy.
Organizational Affiliation: