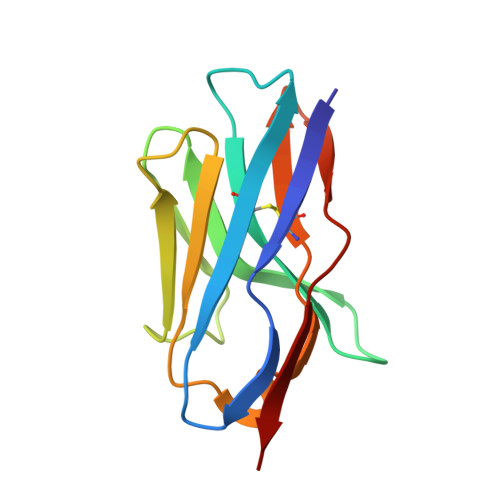
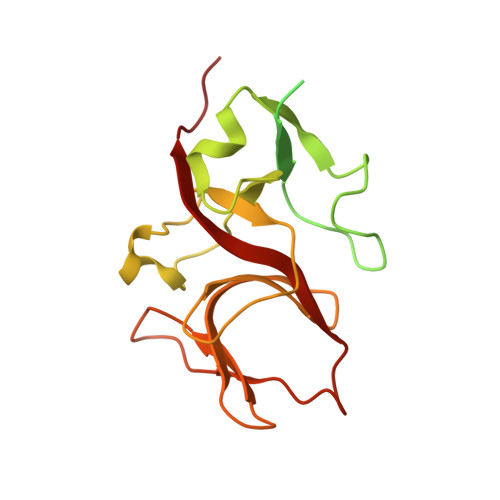
Nanobodies targeting LexA autocleavage disclose a novel suppression strategy of SOS-response pathway.
Maso, L., Vascon, F., Chinellato, M., Goormaghtigh, F., Bellio, P., Campagnaro, E., Van Melderen, L., Ruzzene, M., Pardon, E., Angelini, A., Celenza, G., Steyaert, J., Tondi, D., Cendron, L.(2022) Structure 30: 1479
- PubMed: 36240773 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2022.09.004
- Primary Citation Related Structures:
7B5G, 7OCJ, 7ZRA - PubMed Abstract:
Antimicrobial resistance threatens the eradication of infectious diseases and impairs the efficacy of available therapeutics. The bacterial SOS pathway is a conserved response triggered by genotoxic stresses and represents one of the principal mechanisms that lead to resistance. The RecA recombinase acts as a DNA-damage sensor inducing the autoproteolysis of the transcriptional repressor LexA, thereby derepressing SOS genes that mediate DNA repair, survival to chemotherapy, and hypermutation. The inhibition of such pathway represents a promising strategy for delaying the evolution of antimicrobial resistance. We report the identification, via llama immunization and phage display, of nanobodies that bind LexA with sub-micromolar affinity and block autoproteolysis, repressing SOS response in Escherichia coli. Biophysical characterization of nanobody-LexA complexes revealed that they act by trapping LexA in an inactive conformation and interfering with RecA engagement. Our studies pave the way to the development of new-generation antibiotic adjuvants for the treatment of bacterial infections.
- Dipartimento di Biologia, Università degli Studi di Padova, Via Ugo Bassi 58/b, 35131 Padova, Italy.
Organizational Affiliation: