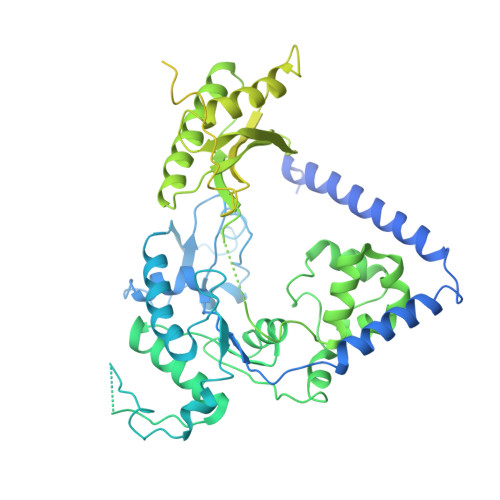
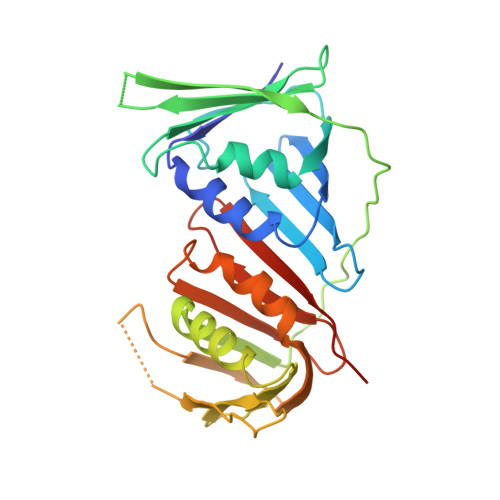
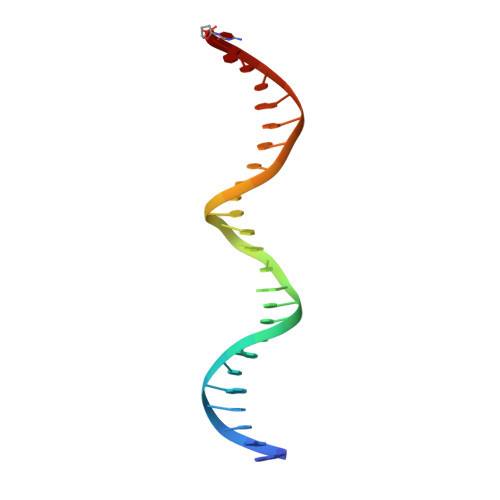
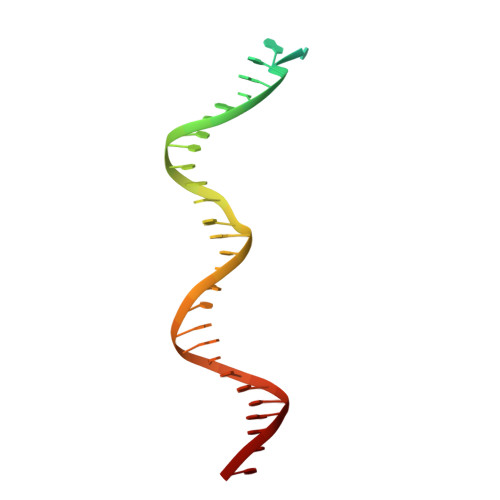
Cryo-EM structure of human Pol kappa bound to DNA and mono-ubiquitylated PCNA.
Lancey, C., Tehseen, M., Bakshi, S., Percival, M., Takahashi, M., Sobhy, M.A., Raducanu, V.S., Blair, K., Muskett, F.W., Ragan, T.J., Crehuet, R., Hamdan, S.M., De Biasio, A.(2021) Nat Commun 12: 6095-6095
- PubMed: 34667155 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-26251-6
- Primary Citation Related Structures:
7NV0, 7NV1 - PubMed Abstract:
Y-family DNA polymerase κ (Pol κ) can replicate damaged DNA templates to rescue stalled replication forks. Access of Pol κ to DNA damage sites is facilitated by its interaction with the processivity clamp PCNA and is regulated by PCNA mono-ubiquitylation. Here, we present cryo-EM reconstructions of human Pol κ bound to DNA, an incoming nucleotide, and wild type or mono-ubiquitylated PCNA (Ub-PCNA). In both reconstructions, the internal PIP-box adjacent to the Pol κ Polymerase-Associated Domain (PAD) docks the catalytic core to one PCNA protomer in an angled orientation, bending the DNA exiting the Pol κ active site through PCNA, while Pol κ C-terminal domain containing two Ubiquitin Binding Zinc Fingers (UBZs) is invisible, in agreement with disorder predictions. The ubiquitin moieties are partly flexible and extend radially away from PCNA, with the ubiquitin at the Pol κ-bound protomer appearing more rigid. Activity assays suggest that, when the internal PIP-box interaction is lost, Pol κ is retained on DNA by a secondary interaction between the UBZs and the ubiquitins flexibly conjugated to PCNA. Our data provide a structural basis for the recruitment of a Y-family TLS polymerase to sites of DNA damage.
- Leicester Institute of Structural & Chemical Biology and Department of Molecular & Cell Biology, University of Leicester, Lancaster Rd, Leicester, LE1 7HB, UK.
Organizational Affiliation: