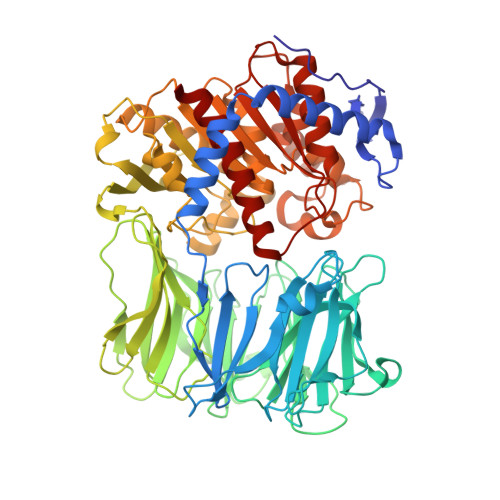
The Crystal Structure of N alpha-p-tosyl-lysyl Chloromethylketone-Bound Oligopeptidase B from Serratia Proteamaculans Revealed a New Type of Inhibitor Binding
Timofeev, V.I., Petrenko, D.E., Agapova, Y.K., Vlaskina, A.V., Karlinsky, D.M., Mikhailova, A.G., Kuranova, I.P., Rakitina, T.V.(2021) Crystals (Basel) 11