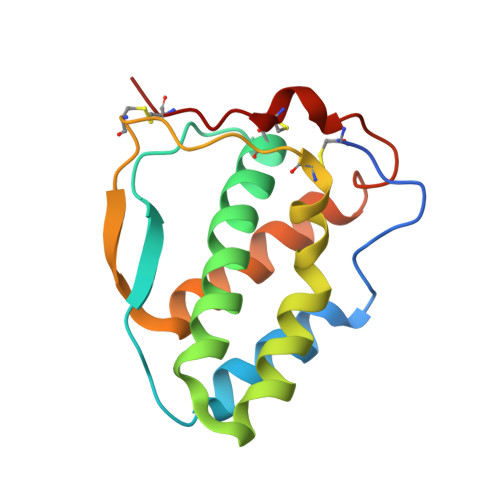
Engineering and crystal structure of a monomeric FLT3 ligand variant.
Pannecoucke, E., Raes, L., Savvides, S.N.(2021) Acta Crystallogr F Struct Biol Commun 77: 121-127
- PubMed: 33830077 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X21003289
- Primary Citation Related Structures:
7NBI - PubMed Abstract:
The overarching paradigm for the activation of class III and V receptor tyrosine kinases (RTKs) prescribes cytokine-mediated dimerization of the receptor ectodomains and homotypic receptor-receptor interactions. However, structural studies have shown that the hematopoietic receptor FLT3, a class III RTK, does not appear to engage in such receptor-receptor contacts, despite its efficient dimerization by dimeric FLT3 ligand (FL). As part of efforts to better understand the intricacies of FLT3 activation, we sought to engineer a monomeric FL. It was found that a Leu27Asp substitution at the dimer interface of the cytokine led to a stable monomeric cytokine (FL L27D ) without abrogation of receptor binding. The crystal structure of FL L27D at 1.65 Å resolution revealed that the introduced point mutation led to shielding of the hydrophobic footprint of the dimerization interface in wild-type FL without affecting the conformation of the FLT3 binding site. Thus, FL L27D can serve as a monomeric FL variant to further interrogate the assembly mechanism of extracellular complexes of FLT3 in physiology and disease.
- Unit for Structural Biology, Department of Biochemistry and Microbiology, Ghent University, Technologiepark-Zwijnaarde 71, 9052 Zwijnaarde, Belgium.
Organizational Affiliation: