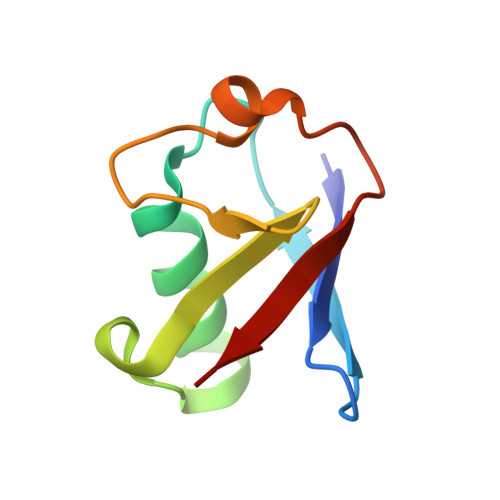
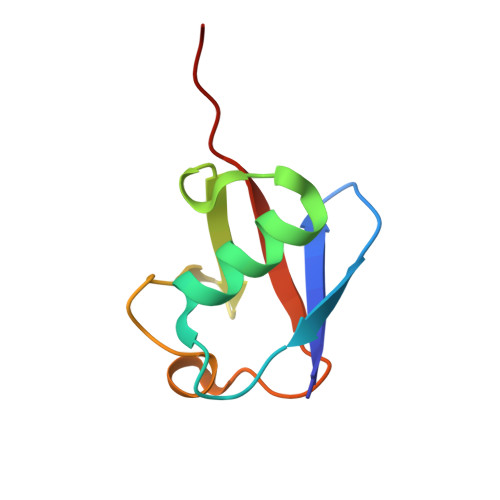
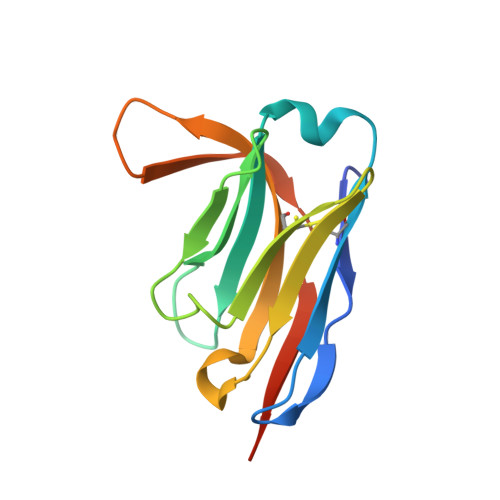
VCP/p97-associated proteins are binders and debranching enzymes of K48-K63-branched ubiquitin chains.
Lange, S.M., McFarland, M.R., Lamoliatte, F., Carroll, T., Krshnan, L., Perez-Rafols, A., Kwasna, D., Shen, L., Wallace, I., Cole, I., Armstrong, L.A., Knebel, A., Johnson, C., De Cesare, V., Kulathu, Y.(2024) Nat Struct Mol Biol
- PubMed: 38977901 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-024-01354-y
- Primary Citation Related Structures:
7NBB, 7NPO, 8A67 - PubMed Abstract:
Branched ubiquitin (Ub) chains constitute a sizable fraction of Ub polymers in human cells. Despite their abundance, our understanding of branched Ub function in cell signaling has been stunted by the absence of accessible methods and tools. Here we identify cellular branched-chain-specific binding proteins and devise approaches to probe K48-K63-branched Ub function. We establish a method to monitor cleavage of linkages within complex Ub chains and unveil ATXN3 and MINDY as debranching enzymes. We engineer a K48-K63 branch-specific nanobody and reveal the molecular basis of its specificity in crystal structures of nanobody-branched Ub chain complexes. Using this nanobody, we detect increased K48-K63-Ub branching following valosin-containing protein (VCP)/p97 inhibition and after DNA damage. Together with our discovery that multiple VCP/p97-associated proteins bind to or debranch K48-K63-linked Ub, these results suggest a function for K48-K63-branched chains in VCP/p97-related processes.
- MRC Protein Phosphorylation and Ubiquitylation Unit, University of Dundee, Dundee, UK. smlange281@gmail.com.
Organizational Affiliation: