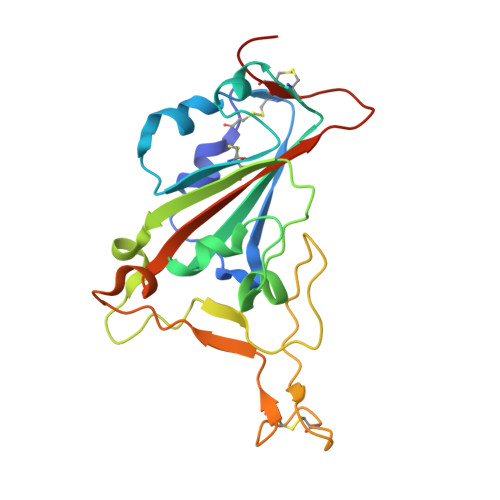
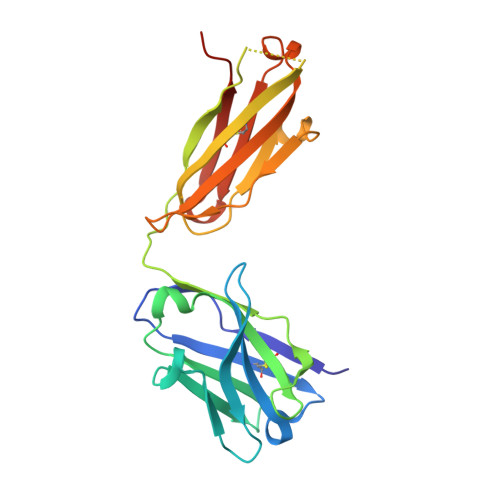
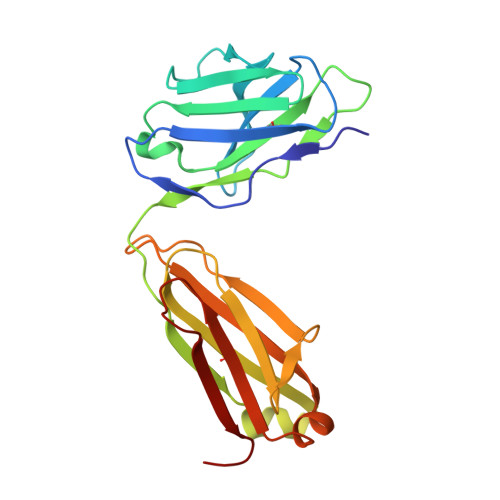
Landscape of human antibody recognition of the SARS-CoV-2 receptor binding domain.
Wheatley, A.K., Pymm, P., Esterbauer, R., Dietrich, M.H., Lee, W.S., Drew, D., Kelly, H.G., Chan, L.J., Mordant, F.L., Black, K.A., Adair, A., Tan, H.X., Juno, J.A., Wragg, K.M., Amarasena, T., Lopez, E., Selva, K.J., Haycroft, E.R., Cooney, J.P., Venugopal, H., Tan, L.L., O Neill, M.T., Allison, C.C., Cromer, D., Davenport, M.P., Bowen, R.A., Chung, A.W., Pellegrini, M., Liddament, M.T., Glukhova, A., Subbarao, K., Kent, S.J., Tham, W.H.(2021) Cell Rep 37: 109822-109822
- PubMed: 34610292 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2021.109822
- Primary Citation Related Structures:
7MZF, 7MZG, 7MZH, 7MZI, 7MZJ, 7MZK, 7MZL, 7MZM, 7MZN, 7RR0 - PubMed Abstract:
Potent neutralizing monoclonal antibodies are one of the few agents currently available to treat COVID-19. SARS-CoV-2 variants of concern (VOCs) that carry multiple mutations in the viral spike protein can exhibit neutralization resistance, potentially affecting the effectiveness of some antibody-based therapeutics. Here, the generation of a diverse panel of 91 human, neutralizing monoclonal antibodies provides an in-depth structural and phenotypic definition of receptor binding domain (RBD) antigenic sites on the viral spike. These RBD antibodies ameliorate SARS-CoV-2 infection in mice and hamster models in a dose-dependent manner and in proportion to in vitro, neutralizing potency. Assessing the effect of mutations in the spike protein on antibody recognition and neutralization highlights both potent single antibodies and stereotypic classes of antibodies that are unaffected by currently circulating VOCs, such as B.1.351 and P.1. These neutralizing monoclonal antibodies and others that bind analogous epitopes represent potentially useful future anti-SARS-CoV-2 therapeutics.
- Department of Microbiology and Immunology, University of Melbourne, the Peter Doherty Institute for Infection and Immunity, Melbourne, VIC 3000, Australia; Australian Research Council Centre for Excellence in Convergent Bio-Nano Science and Technology, University of Melbourne, Melbourne, VIC 3010, Australia.
Organizational Affiliation: