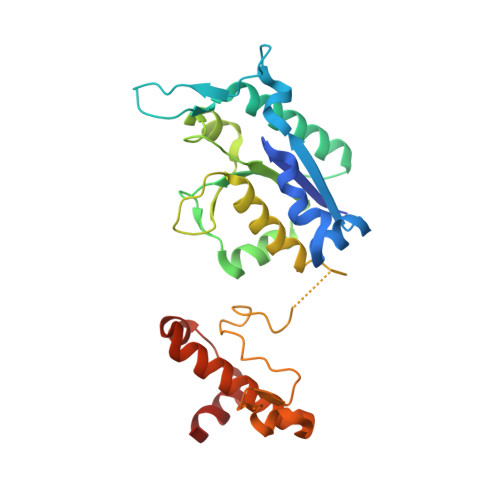
Crystal structure of a tRNA (guanine-N1)-methyltransferase from Acinetobacter baumannii AB5075-UW bound to S-adenosyl homocysteine
Edwards, T.E., Abendroth, J., Horanyi, P.S., Lorimer, D.D., Seattle Structural Genomics Center for Infectious Disease (SSGCID)To be published.