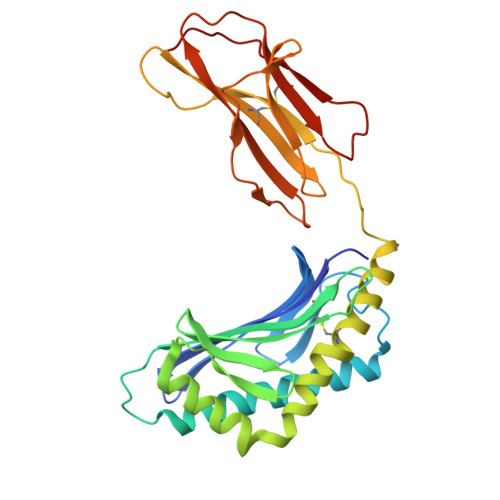
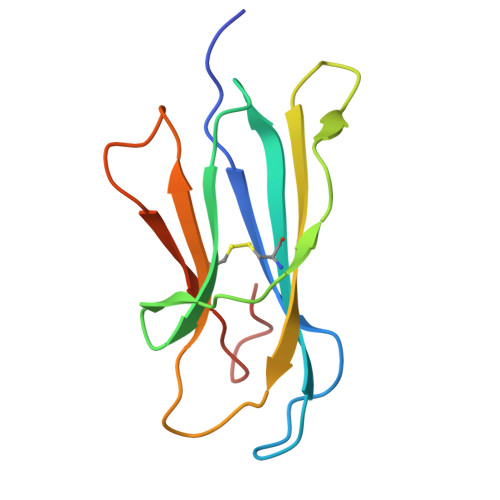
Rational design of a hydrolysis-resistant mycobacterial phosphoglycolipid antigen presented by CD1c to T cells.
Reijneveld, J.F., Marino, L., Cao, T.P., Cheng, T.Y., Dam, D., Shahine, A., Witte, M.D., Filippov, D.V., Suliman, S., van der Marel, G.A., Moody, D.B., Minnaard, A.J., Rossjohn, J., Codee, J.D.C., Van Rhijn, I.(2021) J Biological Chem 297: 101197-101197
- PubMed: 34536421 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2021.101197
- Primary Citation Related Structures:
7MX4, 7MXF, 7MXH - PubMed Abstract:
Whereas proteolytic cleavage is crucial for peptide presentation by classical major histocompatibility complex (MHC) proteins to T cells, glycolipids presented by CD1 molecules are typically presented in an unmodified form. However, the mycobacterial lipid antigen mannosyl-β1-phosphomycoketide (MPM) may be processed through hydrolysis in antigen presenting cells, forming mannose and phosphomycoketide (PM). To further test the hypothesis that some lipid antigens are processed, and to generate antigens that lead to defined epitopes for future tuberculosis vaccines or diagnostic tests, we aimed to create hydrolysis-resistant MPM variants that retain their antigenicity. Here, we designed and tested three different, versatile synthetic strategies to chemically stabilize MPM analogs. Crystallographic studies of CD1c complexes with these three new MPM analogs showed anchoring of the lipid tail and phosphate group that is highly comparable to nature-identical MPM, with considerable conformational flexibility for the mannose head group. MPM-3, a difluoromethylene-modified version of MPM that is resistant to hydrolysis, showed altered recognition by cells, but not by CD1c proteins, supporting the cellular antigen processing hypothesis. Furthermore, the synthetic analogs elicited T cell responses that were cross-reactive with nature-identical MPM, fulfilling important requirements for future clinical use.
- Division of Rheumatology, Inflammation, and Immunity, Brigham and Women's Hospital and Harvard Medical School, Boston, Massachusetts, USA; Department of Infectious Diseases and Immunology, Faculty of Veterinary Medicine, Utrecht University, Utrecht, the Netherlands; Stratingh Institute for Chemistry, University of Groningen, Groningen, the Netherlands.
Organizational Affiliation: