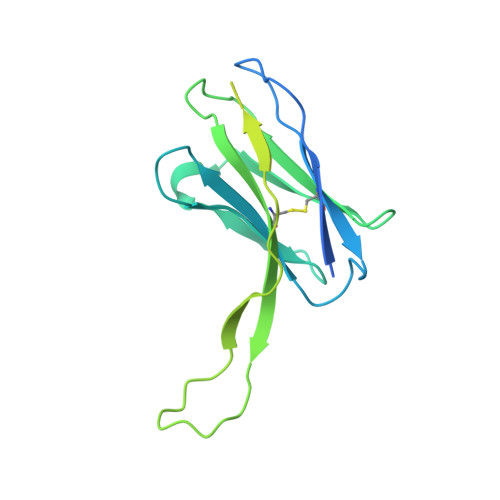
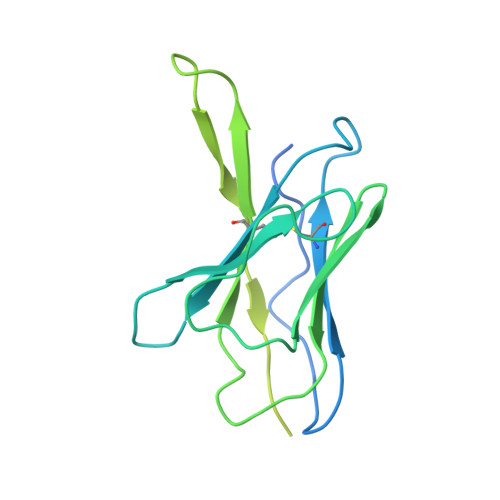
Antibody elicited by HIV-1 immunogen vaccination in macaques displaces Env fusion peptide and destroys a neutralizing epitope.
Abernathy, M.E., Gristick, H.B., Vielmetter, J., Keeffe, J.R., Gnanapragasam, P.N.P., Lee, Y.E., Escolano, A., Gautam, R., Seaman, M.S., Martin, M.A., Nussenzweig, M.C., Bjorkman, P.J.(2021) NPJ Vaccines 6: 126-126
- PubMed: 34697307 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41541-021-00387-4
- Primary Citation Related Structures:
7MXE - PubMed Abstract:
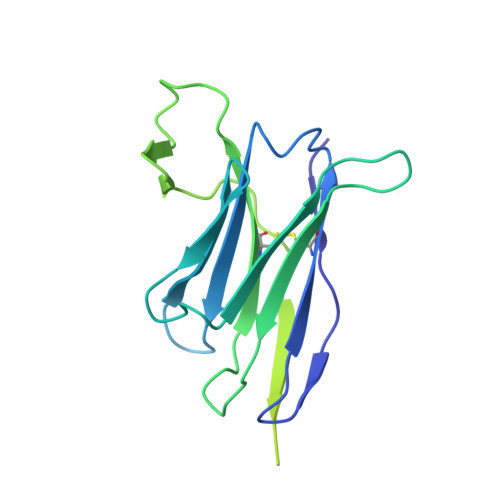
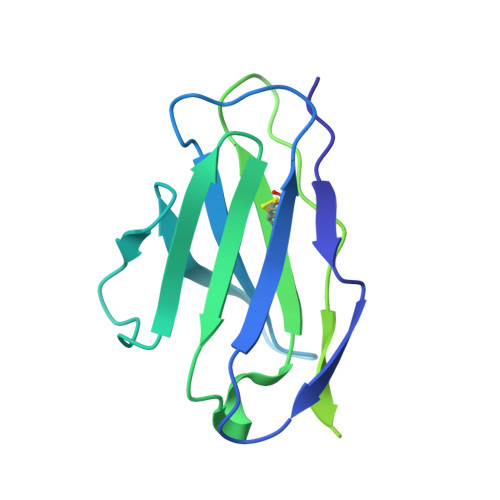
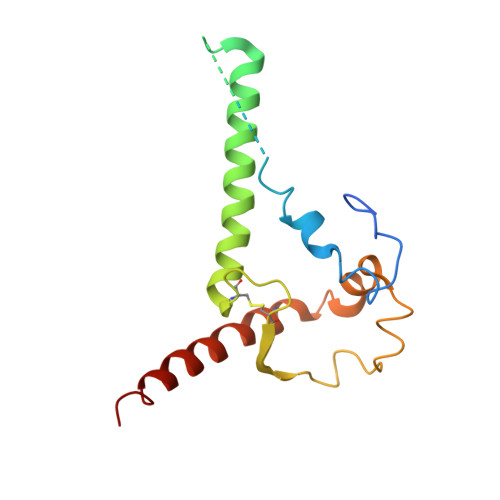
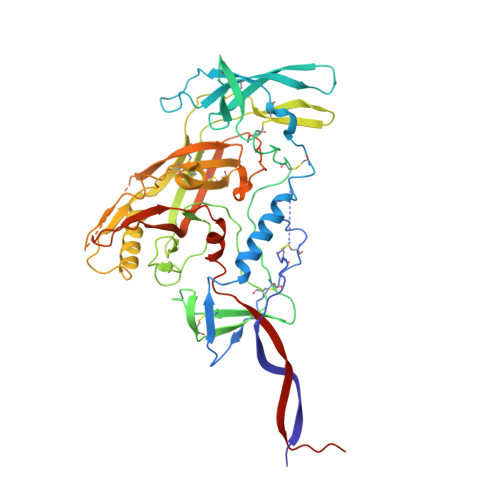
HIV-1 vaccine design aims to develop an immunogen that elicits broadly neutralizing antibodies against a desired epitope, while eliminating responses to off-target regions of HIV-1 Env. We report characterization of Ab1245, an off-target antibody against the Env gp120-gp41 interface, from V3-glycan patch immunogen-primed and boosted macaques. A 3.7 Å cryo-EM structure of an Ab1245-Env complex reveals one Ab1245 Fab binding asymmetrically to Env trimer at the gp120-gp41 interface using its long CDRH3 to mimic regions of gp41. The mimicry includes positioning of a CDRH3 methionine into the gp41 tryptophan clasp, resulting in displacement of the fusion peptide and fusion peptide-proximal region. Despite fusion peptide displacement, Ab1245 is non-neutralizing even at high concentrations, raising the possibility that only two fusion peptides per trimer are required for viral-host membrane fusion. These structural analyses facilitate immunogen design to prevent elicitation of Ab1245-like antibodies that block neutralizing antibodies against the fusion peptide.
- Division of Biology and Biological Engineering, California Institute of Technology, Pasadena, CA, 91125, USA.
Organizational Affiliation: