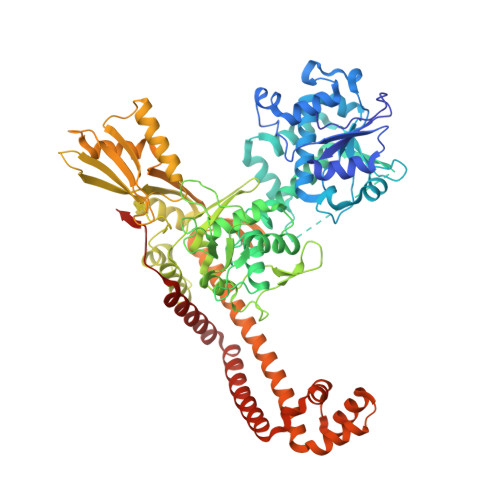
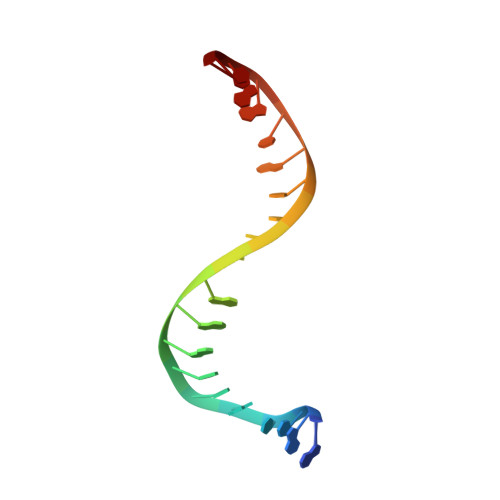
Optimization of TopoIV Potency, ADMET Properties, and hERG Inhibition of 5-Amino-1,3-dioxane-Linked Novel Bacterial Topoisomerase Inhibitors: Identification of a Lead with In Vivo Efficacy against MRSA.
Lu, Y., Vibhute, S., Li, L., Okumu, A., Ratigan, S.C., Nolan, S., Papa, J.L., Mann, C.A., English, A., Chen, A., Seffernick, J.T., Koci, B., Duncan, L.R., Roth, B., Cummings, J.E., Slayden, R.A., Lindert, S., McElroy, C.A., Wozniak, D.J., Yalowich, J., Mitton-Fry, M.J.(2021) J Med Chem 64: 15214-15249
- PubMed: 34614347 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.1c01250
- Primary Citation Related Structures:
7MVS - PubMed Abstract:
Novel bacterial topoisomerase inhibitors (NBTIs) are among the most promising new antibiotics in preclinical/clinical development. We previously reported dioxane-linked NBTIs with potent antistaphylococcal activity and reduced hERG inhibition, a key safety liability. Herein, polarity-focused optimization enabled the delineation of clear structure-property relationships for both microsomal metabolic stability and hERG inhibition, resulting in the identification of lead compound 79 . This molecule demonstrates potent antibacterial activity against diverse Gram-positive pathogens, inhibition of both DNA gyrase and topoisomerase IV, a low frequency of resistance, a favorable in vitro cardiovascular safety profile, and in vivo efficacy in a murine model of methicillin-resistant Staphylococcus aureus infection.
- Division of Medicinal Chemistry and Pharmacognosy, College of Pharmacy, The Ohio State University, Columbus, Ohio 43210, United States.
Organizational Affiliation: