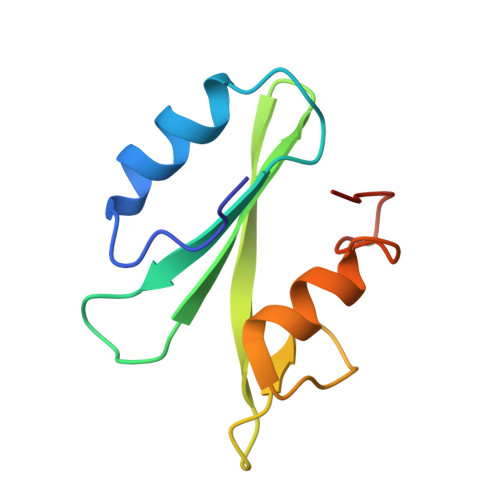
Synthesis and structural characterization of a monocarboxylic inhibitor for GRB2 SH2 domain.
Xiao, T., Sun, L., Zhang, M., Li, Z., Haura, E.B., Schonbrunn, E., Ji, H.(2021) Bioorg Med Chem Lett 51: 128354-128354
- PubMed: 34506932 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bmcl.2021.128354
- Primary Citation Related Structures:
7MPH - PubMed Abstract:
A monocarboxylic inhibitor was designed and synthesized to disrupt the protein-protein interaction (PPI) between GRB2 and phosphotyrosine-containing proteins. Biochemical characterizations show compound 7 binds with the Src homology 2 (SH2) domain of GRB2 and is more potent than EGFR 1068 phosphopeptide 14-mer. X-ray crystallographic studies demonstrate compound 7 occupies the GRB2 binding site for phosphotyrosine-containing sequences and reveal key structural features for GRB2-inhibitor binding. This compound with a -1 formal charge offers a new direction for structural optimization to generate cell-permeable inhibitors for this key protein target of the aberrant Ras-MAPK signaling cascade.
- Drug Discovery Department, H. Lee Moffitt Cancer Center and Research Institute, 12902 Magnolia Drive, Tampa, FL 33612, United States.
Organizational Affiliation: