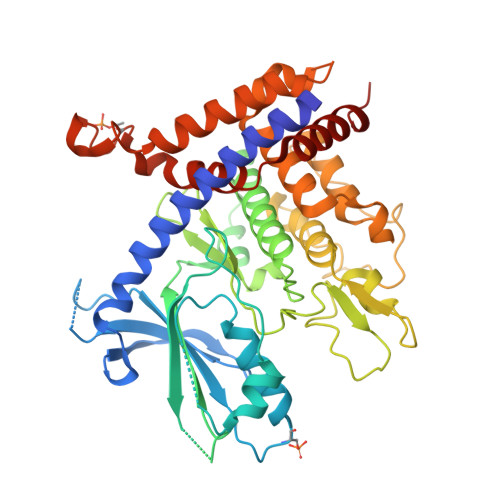
Mechanism of PINK1 activation by autophosphorylation and insights into assembly on the TOM complex.
Rasool, S., Veyron, S., Soya, N., Eldeeb, M.A., Lukacs, G.L., Fon, E.A., Trempe, J.F.(2022) Mol Cell 82: 44
- PubMed: 34875213 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2021.11.012
- Primary Citation Related Structures:
7MP8, 7MP9 - PubMed Abstract:
Mutations in PINK1 cause autosomal-recessive Parkinson's disease. Mitochondrial damage results in PINK1 import arrest on the translocase of the outer mitochondrial membrane (TOM) complex, resulting in the activation of its ubiquitin kinase activity by autophosphorylation and initiation of Parkin-dependent mitochondrial clearance. Herein, we report crystal structures of the entire cytosolic domain of insect PINK1. Our structures reveal a dimeric autophosphorylation complex targeting phosphorylation at the invariant Ser205 (human Ser228). The dimer interface requires insert 2, which is unique to PINK1. The structures also reveal how an N-terminal helix binds to the C-terminal extension and provide insights into stabilization of PINK1 on the core TOM complex.
- Department of Pharmacology & Therapeutics and Centre de Recherche en Biologie Structurale, McGill University, Montréal, QC, Canada.
Organizational Affiliation: