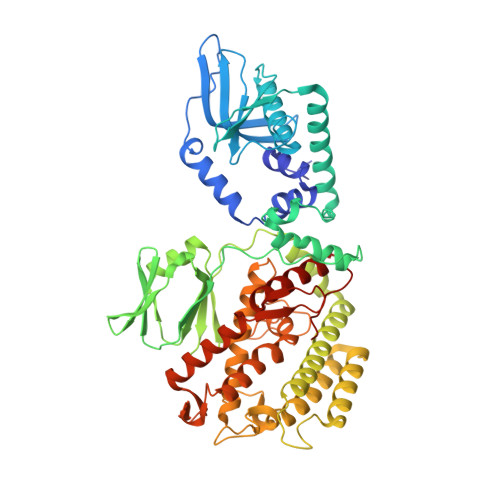
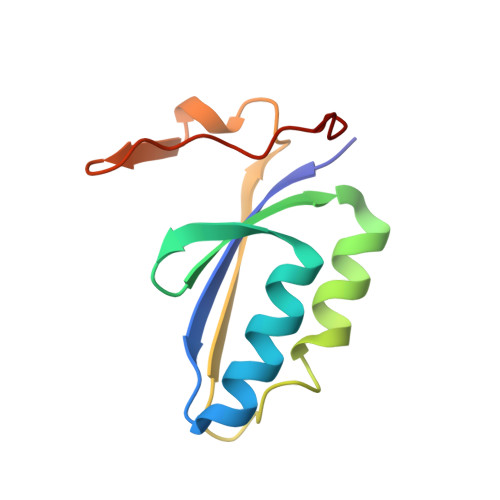
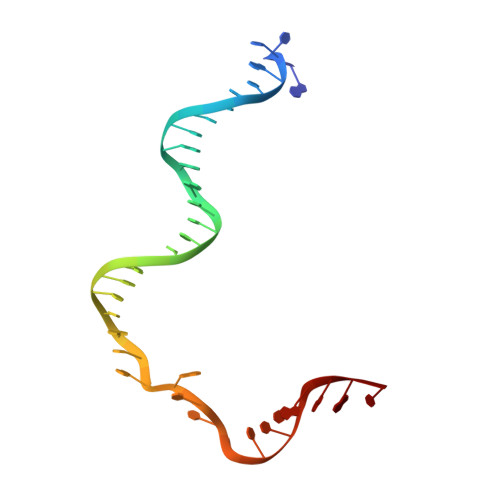
Mechanism for Cas4-assisted directional spacer acquisition in CRISPR-Cas.
Hu, C., Almendros, C., Nam, K.H., Costa, A.R., Vink, J.N.A., Haagsma, A.C., Bagde, S.R., Brouns, S.J.J., Ke, A.(2021) Nature 598: 515-520
- PubMed: 34588691 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-021-03951-z
- Primary Citation Related Structures:
7MI4, 7MI5, 7MI9, 7MIB, 7MID - PubMed Abstract:
Prokaryotes adapt to challenges from mobile genetic elements by integrating spacers derived from foreign DNA in the CRISPR array 1 . Spacer insertion is carried out by the Cas1-Cas2 integrase complex 2-4 . A substantial fraction of CRISPR-Cas systems use a Fe-S cluster containing Cas4 nuclease to ensure that spacers are acquired from DNA flanked by a protospacer adjacent motif (PAM) 5,6 and inserted into the CRISPR array unidirectionally, so that the transcribed CRISPR RNA can guide target searching in a PAM-dependent manner. Here we provide a high-resolution mechanistic explanation for the Cas4-assisted PAM selection, spacer biogenesis and directional integration by type I-G CRISPR in Geobacter sulfurreducens, in which Cas4 is naturally fused with Cas1, forming Cas4/Cas1. During biogenesis, only DNA duplexes possessing a PAM-embedded 3'-overhang trigger Cas4/Cas1-Cas2 assembly. During this process, the PAM overhang is specifically recognized and sequestered, but is not cleaved by Cas4. This 'molecular constipation' prevents the PAM-side prespacer from participating in integration. Lacking such sequestration, the non-PAM overhang is trimmed by host nucleases and integrated to the leader-side CRISPR repeat. Half-integration subsequently triggers PAM cleavage and Cas4 dissociation, allowing spacer-side integration. Overall, the intricate molecular interaction between Cas4 and Cas1-Cas2 selects PAM-containing prespacers for integration and couples the timing of PAM processing with the stepwise integration to establish directionality.
- Department of Molecular Biology and Genetics, Cornell University, Ithaca, NY, USA.
Organizational Affiliation: