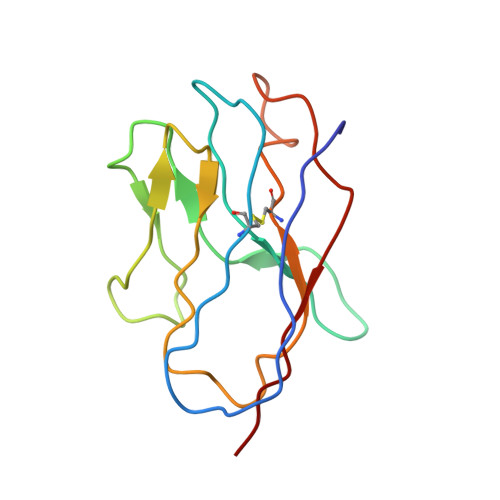
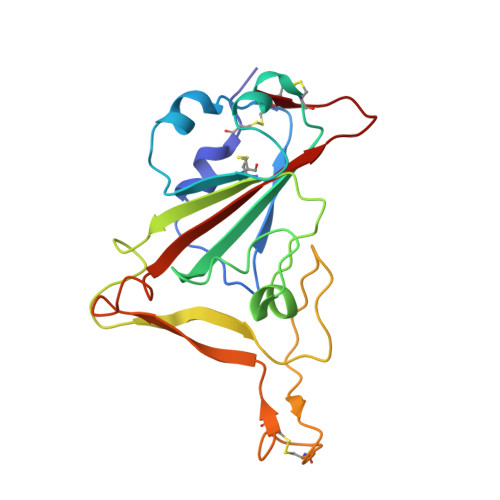
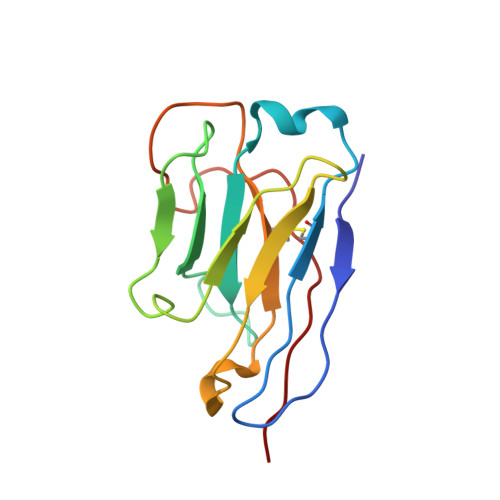
Potent neutralizing nanobodies resist convergent circulating variants of SARS-CoV-2 by targeting diverse and conserved epitopes
Sun, D., Sang, Z., Kim, Y.J., Xiang, Y., Cohen, T., Belford, A.K., Huet, A., Conway, J.F., Sun, J., Taylor, D.J., Schneidman-Duhovny, D., Zhang, C., Huang, W., Shi, Y.(2021) Nat Commun 12: 4676