Pan-ebolavirus protective therapy by two multifunctional human antibodies.
Gilchuk, P., Murin, C.D., Cross, R.W., Ilinykh, P.A., Huang, K., Kuzmina, N., Borisevich, V., Agans, K.N., Geisbert, J.B., Zost, S.J., Nargi, R.S., Sutton, R.E., Suryadevara, N., Bombardi, R.G., Carnahan, R.H., Bukreyev, A., Geisbert, T.W., Ward, A.B., Crowe Jr., J.E.(2021) Cell 184: 5593-5607.e18
- PubMed: 34715022 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2021.09.035
- Primary Citation Related Structures:
7M8L - PubMed Abstract:
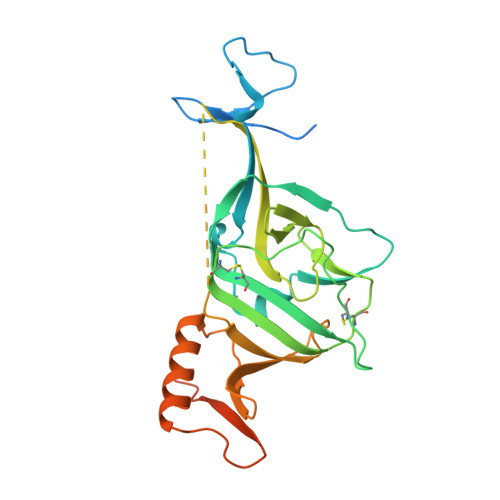
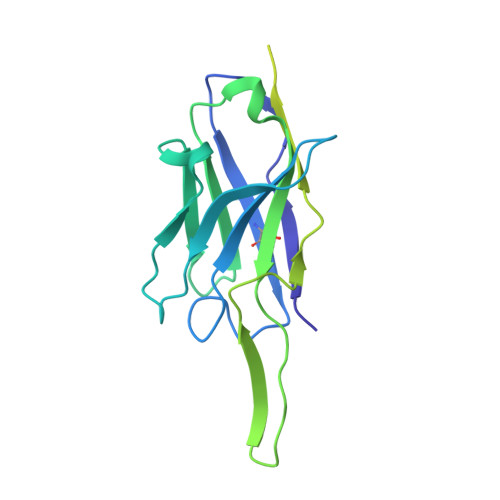
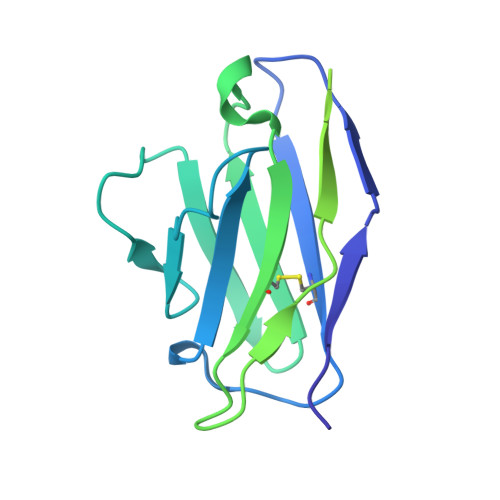
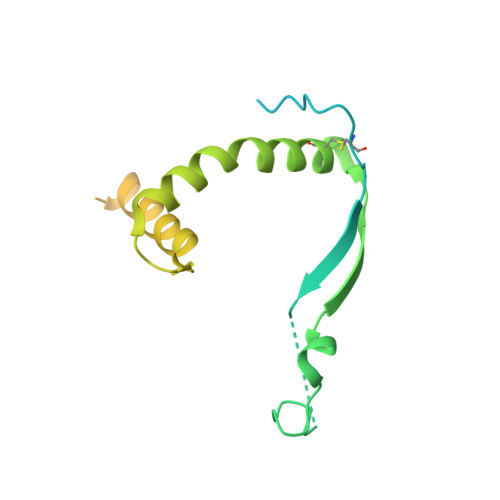
Ebolaviruses cause a severe and often fatal illness with the potential for global spread. Monoclonal antibody-based treatments that have become available recently have a narrow therapeutic spectrum and are ineffective against ebolaviruses other than Ebola virus (EBOV), including medically important Bundibugyo (BDBV) and Sudan (SUDV) viruses. Here, we report the development of a therapeutic cocktail comprising two broadly neutralizing human antibodies, rEBOV-515 and rEBOV-442, that recognize non-overlapping sites on the ebolavirus glycoprotein (GP). Antibodies in the cocktail exhibited synergistic neutralizing activity, resisted viral escape, and possessed differing requirements for their Fc-regions for optimal in vivo activities. The cocktail protected non-human primates from ebolavirus disease caused by EBOV, BDBV, or SUDV with high therapeutic effectiveness. High-resolution structures of the cocktail antibodies in complex with GP revealed the molecular determinants for neutralization breadth and potency. This study provides advanced preclinical data to support clinical development of this cocktail for pan-ebolavirus therapy.
- Vanderbilt Vaccine Center, Vanderbilt University Medical Center, Nashville, TN 37232, USA.
Organizational Affiliation: